Short-term metabolic adjustments in Plasmodium falciparum counter hypoxanthine deprivation at the expense of long-term viability
- PMID: 30890151
- PMCID: PMC6423861
- DOI: 10.1186/s12936-019-2720-3
Short-term metabolic adjustments in Plasmodium falciparum counter hypoxanthine deprivation at the expense of long-term viability
Abstract
Background: The malarial parasite Plasmodium falciparum is an auxotroph for purines, which are required for nucleic acid synthesis during the intra-erythrocytic developmental cycle (IDC) of the parasite. The capabilities of the parasite and extent to which it can use compensatory mechanisms to adapt to purine deprivation were studied by examining changes in its metabolism under sub-optimal concentrations of hypoxanthine, the primary precursor utilized by the parasite for purine-based nucleic acid synthesis.
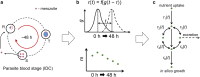
Methods: The concentration of hypoxanthine that caused a moderate growth defect over the course of one IDC was determined. At this concentration of hypoxanthine (0.5 μM), transcriptomic and metabolomic data were collected during one IDC at multiple time points. These data were integrated with a metabolic network model of the parasite embedded in a red blood cell (RBC) to interpret the metabolic adaptation of P. falciparum to hypoxanthine deprivation.
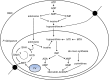
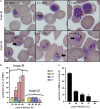
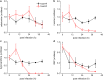
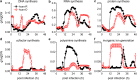
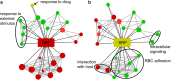
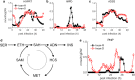
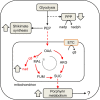
Results: At a hypoxanthine concentration of 0.5 μM, vacuole-like structures in the cytosol of many P. falciparum parasites were observed after the 24-h midpoint of the IDC. Parasites grown under these conditions experienced a slowdown in the progression of the IDC. After 72 h of deprivation, the parasite growth could not be recovered despite supplementation with 90 µM hypoxanthine. Simulations of P. falciparum metabolism suggested that alterations in ubiquinone, isoprenoid, shikimate, and mitochondrial metabolism occurred before the appearance of these vacuole-like structures. Alterations were found in metabolic reactions associated with fatty acid synthesis, the pentose phosphate pathway, methionine metabolism, and coenzyme A synthesis in the latter half of the IDC. Furthermore, gene set enrichment analysis revealed that P. falciparum activated genes associated with rosette formation, Maurer's cleft and protein export under two different nutrient-deprivation conditions (hypoxanthine and isoleucine).
Conclusions: The metabolic network analysis presented here suggests that P. falciparum invokes specific purine-recycling pathways to compensate for hypoxanthine deprivation and maintains a hypoxanthine pool for purine-based nucleic acid synthesis. However, this compensatory mechanism is not sufficient to maintain long-term viability of the parasite. Although P. falciparum can complete a full IDC in low hypoxanthine conditions, subsequent cycles are disrupted.
Keywords: Gene set enrichment analysis; Metabolic network model; Metabolome; Plasmodium falciparum; Purine deprivation; Stress response pathways; Transcriptome.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Metabolic Survival Adaptations of Plasmodium falciparum Exposed to Sublethal Doses of Fosmidomycin.Antimicrob Agents Chemother. 2021 Mar 18;65(4):e02392-20. doi: 10.1128/AAC.02392-20. Print 2021 Mar 18. Antimicrob Agents Chemother. 2021. PMID: 33495219 Free PMC article.
-
Modeling metabolism and stage-specific growth of Plasmodium falciparum HB3 during the intraerythrocytic developmental cycle.Mol Biosyst. 2014 Oct;10(10):2526-37. doi: 10.1039/c4mb00115j. Mol Biosyst. 2014. PMID: 25001103
-
Transport of purines and purine salvage pathway inhibitors by the Plasmodium falciparum equilibrative nucleoside transporter PfENT1.Mol Biochem Parasitol. 2010 Jan;169(1):40-9. doi: 10.1016/j.molbiopara.2009.10.001. Epub 2009 Oct 8. Mol Biochem Parasitol. 2010. PMID: 19818813 Free PMC article.
-
Metabolic changes of the malaria parasite during the transition from the human to the mosquito host.Annu Rev Microbiol. 1998;52:561-90. doi: 10.1146/annurev.micro.52.1.561. Annu Rev Microbiol. 1998. PMID: 9891808 Review.
-
Using metabolomics to dissect host-parasite interactions.Curr Opin Microbiol. 2016 Aug;32:59-65. doi: 10.1016/j.mib.2016.04.019. Epub 2016 May 17. Curr Opin Microbiol. 2016. PMID: 27200489 Review.
Cited by
-
Metabolic adjustments of blood-stage Plasmodium falciparum in response to sublethal pyrazoleamide exposure.Sci Rep. 2022 Jan 21;12(1):1167. doi: 10.1038/s41598-022-04985-7. Sci Rep. 2022. PMID: 35064153 Free PMC article.
-
Metabolic changes accompanying the loss of fumarate hydratase and malate-quinone oxidoreductase in the asexual blood stage of Plasmodium falciparum.J Biol Chem. 2022 May;298(5):101897. doi: 10.1016/j.jbc.2022.101897. Epub 2022 Apr 6. J Biol Chem. 2022. PMID: 35398098 Free PMC article.
-
Inter-study and time-dependent variability of metabolite abundance in cultured red blood cells.Malar J. 2021 Jul 2;20(1):299. doi: 10.1186/s12936-021-03780-5. Malar J. 2021. PMID: 34215262 Free PMC article.
-
Metabolic Survival Adaptations of Plasmodium falciparum Exposed to Sublethal Doses of Fosmidomycin.Antimicrob Agents Chemother. 2021 Mar 18;65(4):e02392-20. doi: 10.1128/AAC.02392-20. Print 2021 Mar 18. Antimicrob Agents Chemother. 2021. PMID: 33495219 Free PMC article.
-
A mevalonate bypass system facilitates elucidation of plastid biology in malaria parasites.PLoS Pathog. 2020 Feb 14;16(2):e1008316. doi: 10.1371/journal.ppat.1008316. eCollection 2020 Feb. PLoS Pathog. 2020. PMID: 32059044 Free PMC article.
References
-
- WHO . World malaria report 2017. Geneva: World Health Organization; 2018.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

