Species-specific enhancement of enterohemorrhagic E. coli pathogenesis mediated by microbiome metabolites
- PMID: 30890187
- PMCID: PMC6425591
- DOI: 10.1186/s40168-019-0650-5
Species-specific enhancement of enterohemorrhagic E. coli pathogenesis mediated by microbiome metabolites
Abstract
Background: Species-specific differences in tolerance to infection are exemplified by the high susceptibility of humans to enterohemorrhagic Escherichia coli (EHEC) infection, whereas mice are relatively resistant to this pathogen. This intrinsic species-specific difference in EHEC infection limits the translation of murine research to human. Furthermore, studying the mechanisms underlying this differential susceptibility is a difficult problem due to complex in vivo interactions between the host, pathogen, and disparate commensal microbial communities.
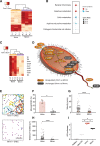
Results: We utilize organ-on-a-chip (Organ Chip) microfluidic culture technology to model damage of the human colonic epithelium induced by EHEC infection, and show that epithelial injury is greater when exposed to metabolites derived from the human gut microbiome compared to mouse. Using a multi-omics approach, we discovered four human microbiome metabolites-4-methyl benzoic acid, 3,4-dimethylbenzoic acid, hexanoic acid, and heptanoic acid-that are sufficient to mediate this effect. The active human microbiome metabolites preferentially induce expression of flagellin, a bacterial protein associated with motility of EHEC and increased epithelial injury. Thus, the decreased tolerance to infection observed in humans versus other species may be due in part to the presence of compounds produced by the human intestinal microbiome that actively promote bacterial pathogenicity.
Conclusion: Organ-on-chip technology allowed the identification of specific human microbiome metabolites modulating EHEC pathogenesis. These identified metabolites are sufficient to increase susceptibility to EHEC in our human Colon Chip model and they contribute to species-specific tolerance. This work suggests that higher concentrations of these metabolites could be the reason for higher susceptibility to EHEC infection in certain human populations, such as children. Furthermore, this research lays the foundation for therapeutic-modulation of microbe products in order to prevent and treat human bacterial infection.
Conflict of interest statement
Ethics approval and consent to participate
Human colonic resections were obtained anonymously from the Department of Pathology at Massachusetts General Hospital under an existing Institutional Review Board approved protocol (#2015P001859). Endoscopic biopsies were collected from de-identified patients from Boston Children’s Hospital. Informed consent and developmentally appropriate assent were obtained at Boston Children’s Hospital from the donors’ guardian and the donor, respectively. All methods were carried out in accordance with the Institutional Review Board of Boston Children’s Hospital (Protocol number IRB-P00000529) approval.
Consent for publication
Not applicable.
Competing interests
D.E.I. holds equity in Emulate, Inc., consults to the company, and chairs its scientific advisory board; he also is an inventor on relevant patents.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



Similar articles
-
Interactions between Enterohemorrhagic Escherichia coli (EHEC) and Gut Commensals at the Interface of Human Colonoids.mBio. 2022 Jun 28;13(3):e0132122. doi: 10.1128/mbio.01321-22. Epub 2022 May 31. mBio. 2022. PMID: 35638758 Free PMC article.
-
The Canonical Long-Chain Fatty Acid Sensing Machinery Processes Arachidonic Acid To Inhibit Virulence in Enterohemorrhagic Escherichia coli.mBio. 2021 Jan 19;12(1):e03247-20. doi: 10.1128/mBio.03247-20. mBio. 2021. PMID: 33468701 Free PMC article.
-
Enterohemorrhagic Escherichia coli (EHEC) disrupts intestinal barrier integrity in translational canine stem cell-derived monolayers.Microbiol Spectr. 2024 Oct 3;12(10):e0096124. doi: 10.1128/spectrum.00961-24. Epub 2024 Aug 20. Microbiol Spectr. 2024. PMID: 39162490 Free PMC article.
-
Zoonotic enterohemorrhagic Escherichia coli: A One Health perspective.ILAR J. 2010;51(3):221-32. doi: 10.1093/ilar.51.3.221. ILAR J. 2010. PMID: 21131723 Review.
-
The role of lymphostatin/EHEC factor for adherence-1 in the pathogenesis of gram negative infection.Toxins (Basel). 2010 May;2(5):954-62. doi: 10.3390/toxins2050954. Epub 2010 May 5. Toxins (Basel). 2010. PMID: 22069619 Free PMC article. Review.
Cited by
-
InVitro Models of Intestine Innate Immunity.Trends Biotechnol. 2021 Mar;39(3):274-285. doi: 10.1016/j.tibtech.2020.07.009. Epub 2020 Aug 24. Trends Biotechnol. 2021. PMID: 32854949 Free PMC article. Review.
-
Human intestinal models to study interactions between intestine and microbes.Open Biol. 2020 Oct;10(10):200199. doi: 10.1098/rsob.200199. Epub 2020 Oct 21. Open Biol. 2020. PMID: 33081633 Free PMC article.
-
Bio-Microfabrication of 2D and 3D Biomimetic Gut-on-a-Chip.Micromachines (Basel). 2023 Sep 4;14(9):1736. doi: 10.3390/mi14091736. Micromachines (Basel). 2023. PMID: 37763899 Free PMC article. Review.
-
Critical Considerations for the Design of Multi-Organ Microphysiological Systems (MPS).Front Cell Dev Biol. 2021 Sep 9;9:721338. doi: 10.3389/fcell.2021.721338. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34568333 Free PMC article. Review.
-
Organs-on-chips: into the next decade.Nat Rev Drug Discov. 2021 May;20(5):345-361. doi: 10.1038/s41573-020-0079-3. Epub 2020 Sep 10. Nat Rev Drug Discov. 2021. PMID: 32913334 Review.
References
-
- Cheng Y-L, Song L-Q, Huang Y-M, Xiong Y-W, Zhang X-A, Sun H, et al. Effect of enterohaemorrhagic Escherichia coli O157:H7-specific enterohaemolysin on interleukin-1β production differs between human and mouse macrophages due to the different sensitivity of NLRP3 activation. Immunology. 2015;145:258–267. doi: 10.1111/imm.12442. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

