Affinity depletion versus relative protein enrichment: a side-by-side comparison of two major strategies for increasing human cerebrospinal fluid proteome coverage
- PMID: 30890900
- PMCID: PMC6390343
- DOI: 10.1186/s12014-019-9229-1
Affinity depletion versus relative protein enrichment: a side-by-side comparison of two major strategies for increasing human cerebrospinal fluid proteome coverage
Abstract
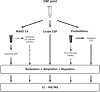
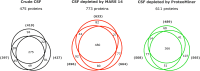
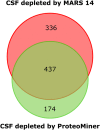
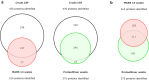
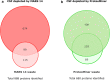
Cerebrospinal fluid (CSF) is in direct contact with the central nervous system. This makes human CSF an attractive source of potential biomarkers for neurologic diseases. Similarly to blood plasma, proteomic analysis of CSF is complicated by a high dynamic range of individual protein concentrations and by the presence of several highly abundant proteins. To deal with the abundant human CSF proteins, methods developed for blood plasma/serum are routinely used. Multiple affinity removal systems and protein enrichment of less abundant proteins using a combinatorial peptide ligand library are among the most frequent approaches. However, their relative impact on CSF proteome coverage has never been evaluated side-by-side in a single study. Therefore, we explored the effect of CSF depletion using MARS 14 cartridge and ProteoMiner ligand library on the number of CSF proteins identified in subsequent LC-MS/MS analysis. LC-MS/MS analysis of crude (non-treated) CSF provided roughly 500 identified proteins. Depletion of CSF by MARS 14 cartridge increased the number of identifications to nearly 800, while treatment of CSF using ProteoMiner enabled identification of 600 proteins. To explore the potential losses of CSF proteins during the depletion process, we also analyzed the "waste" fractions generated by both methods, i.e., proteins retained by the MARS 14 cartridge, and the molecules present in the flow-through fraction from ProteoMiner. More than 250 proteins were bound to MARS 14 cartridge, 100 of those were not identified in the corresponding depleted CSF. Similarly, analysis of the waste fraction in ProteoMiner workflow provided almost 70 unique proteins not found in the CSF depleted by the ligand library. Both depletion strategies significantly increased the number of identified CSF proteins compared to crude CSF. However, MARS 14 depletion provided a markedly higher number of identified proteins (773) compared to ProteoMiner (611). Further, we showed that CSF proteins are lost due to co-depletion (MARS 14) or exclusion (ProteoMiner) during the depletion process. This suggests that the routinely discarded "waste" fractions contain proteins of potential interest and should be included in CSF biomarker studies.
Keywords: Biomarkers; CNS; Cerebrospinal fluid; Depletion; LC–MS/MS; Mass spectrometry.
Figures
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
