PTPN14 degradation by high-risk human papillomavirus E7 limits keratinocyte differentiation and contributes to HPV-mediated oncogenesis
- PMID: 30894485
- PMCID: PMC6452706
- DOI: 10.1073/pnas.1819534116
PTPN14 degradation by high-risk human papillomavirus E7 limits keratinocyte differentiation and contributes to HPV-mediated oncogenesis
Abstract
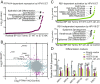
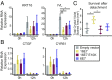
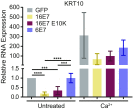
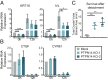
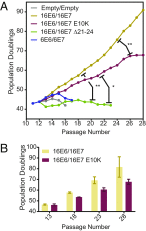
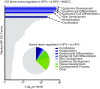
High-risk human papillomavirus (HPV) E7 proteins enable oncogenic transformation of HPV-infected cells by inactivating host cellular proteins. High-risk but not low-risk HPV E7 target PTPN14 for proteolytic degradation, suggesting that PTPN14 degradation may be related to their oncogenic activity. HPV infects human keratinocytes but the role of PTPN14 in keratinocytes and the consequences of PTPN14 degradation are unknown. Using an HPV16 E7 variant that can inactivate retinoblastoma tumor suppressor (RB1) but cannot degrade PTPN14, we found that high-risk HPV E7-mediated PTPN14 degradation impairs keratinocyte differentiation. Deletion of PTPN14 from primary human keratinocytes decreased keratinocyte differentiation gene expression. Related to oncogenic transformation, both HPV16 E7-mediated PTPN14 degradation and PTPN14 deletion promoted keratinocyte survival following detachment from a substrate. PTPN14 degradation contributed to high-risk HPV E6/E7-mediated immortalization of primary keratinocytes and HPV+ but not HPV- cancers exhibit a gene-expression signature consistent with PTPN14 inactivation. We find that PTPN14 degradation impairs keratinocyte differentiation and propose that this contributes to high-risk HPV E7-mediated oncogenic activity independent of RB1 inactivation.
Keywords: HPV; PTPN14; carcinogenesis; differentiation; papillomavirus.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Graham SV. The human papillomavirus replication cycle, and its links to cancer progression: A comprehensive review. Clin Sci (Lond) 2017;131:2201–2221. - PubMed
-
- Muñoz N, et al. International Agency for Research on Cancer Multicenter Cervical Cancer Study Group Epidemiologic classification of human papillomavirus types associated with cervical cancer. N Engl J Med. 2003;348:518–527. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

