miR-363 acts as a tumor suppressor in osteosarcoma cells by inhibiting PDZD2
- PMID: 30896877
- PMCID: PMC6448123
- DOI: 10.3892/or.2019.7078
miR-363 acts as a tumor suppressor in osteosarcoma cells by inhibiting PDZD2
Abstract
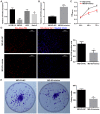
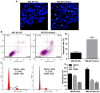
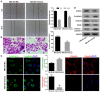
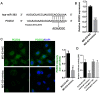
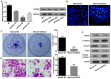
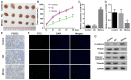
PDZ domain containing 2 (PDZD2) is a multi-PDZ domain protein that promotes the proliferation of insulinoma cells, and is upregulated during prostate tumorigenesis. However, the function of PDZD2 in other cancers, including osteosarcoma (OS), remains unclear. Dysregulation of microRNAs (miRNAs) contributes to tumor initiation, proliferation and metastasis, via the regulation of their target genes. The present study investigated the functions of miR-363 and PDZD2 in MG-63 OS cells. The results revealed that MG-63 cells contained low levels of miR-363, and that overexpression of miR-363 in MG-63 cells significantly inhibited the vitality, proliferation, and colony formation ability of the cells, but promoted their apoptosis and G1/S arrest by regulating proliferating cell nuclear antigen (PCNA) and caspase-3 expression. Additionally, miR-363 impaired the migration and invasion of MG-63 cells by regulating the epithelial-mesenchymal transition (EMT) phenotype. Notably, a bioinformatics analysis and luciferase reporter assay indicated that PDZD2 was a direct target of miR-363. miR-363 overexpression reduced PDZD2 protein levels and knockdown of PDZD2 suppressed the colony formation, migration and invasion of MG-63 cells, but promoted their apoptosis by regulating expression of PCNA, caspase-3, and the EMT phenotype. In vivo studies further confirmed that miR-363 functioned as tumor suppressor, by inhibiting tumor growth, promoting cell apoptosis, and reducing PDZD2 and PCNA levels and the prevalence of the EMT phenotype in tumor tissues. The present data demonstrated that downregulation of the tumor suppressor miR-363 may be involved in the development of osteosarcoma via regulation of PDZD2.
Figures
References
-
- Geller DS, Gorlick R. Osteosarcoma: A review of diagnosis, management, and treatment strategies. Clin Adv Hematol Oncol. 2010;8:705–718. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

