Vortex- and Centrifugation-Free Extraction of HIV-1 RNA
- PMID: 30911908
- PMCID: PMC11289783
- DOI: 10.1007/s40291-019-00394-1
Vortex- and Centrifugation-Free Extraction of HIV-1 RNA
Abstract
Background and objective: HIV viral load measurements play a critical role in monitoring disease progression in those who are on antiretroviral treatment. In order to obtain an accurate measurement, rapid sample preparation techniques are required. There is an unmet need for HIV extraction instruments in resource-limited settings, where HIV prevalence is high. Therefore, the objective of our study was to develop a three-dimensional (3D) microfluidic system to extract HIV-1 RNA with minimal electricity and without complex laboratory instruments.
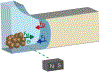
Methods: A 3D microfluidic system was designed in which magnetic beads bound with nucleic acids move through immiscible oil-water interfaces to separate HIV-1 RNA from the sample. Polymerase chain reaction (PCR) amplification was used to quantify the total amount of HIV-1 RNA extracted as we optimized the system through chip design, bead type, carry-over volume, carrier RNA concentration, and elution buffer temperature. Additionally, the extraction efficiency of the 3D microfluidic system was evaluated by comparing with a Qiagen EZ1 Advanced XL instrument using 20 HIV-1-positive plasma samples.
Results: Our method has near-perfect (100%) extraction efficiency in spiked serum samples with as little as 50 copies/mL starting sample. Furthermore, we report carry-over volumes of 0.31% ± 0.006% of total sample volume. Using the EZ1 Advanced XL as a gold standard, the average percentage HIV-1 RNA extracted using the microchip was observed to be 65.4% ± 24.6%.
Conclusions: From a clinical perspective, the success of our method opens up its possible use in diagnostic tests for HIV in the remote areas where access to vortexes and centrifuges is not available. Here we present a proof-of-concept device which, with further development, could be used for sample preparation at the point of care.
Conflict of interest statement
Compliance with Ethical Standards
Figures





References
-
- Deng MY, Wang H, Ward GB, Beckham TR, McKenna TS. Comparison of six RNA extraction methods for the detection of classical swine fever virus by real-time and conventional reverse transcription-PCR. J Vet Diagn Investig. 2005;17:574–8. - PubMed
-
- Monleau M, Montavon C, Laurent C, Segondy M, Montes B, Delaporte E, et al. Evaluation of different RNA extraction methods and storage conditions of dried plasma or blood spots for human immunodeficiency virus type 1 RNA quantification and PCR amplification for drug resistance testing. J Clin Microbiol. 2009;47:1107–18. - PMC - PubMed
-
- Rahman MM, Elaissari A. Nucleic acid sample preparation for in vitro molecular diagnosis: from conventional techniques to biotechnology. Drug Discov Today. 2012;17:1199–207. - PubMed
-
- UNAIDS. How AIDS changed everything—MDG6: 15 years, 15 lessons of hope from the AIDS response. 2015. 10.1007/s13398-014-0173-7.2. http://www.unaids.org/en/resources/documents/2015/MDG6_15years-15lessons.... Accessed 6 Mar 2019. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

