Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution
- PMID: 30923225
- PMCID: PMC6927209
- DOI: 10.1126/science.aaw1219
Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution
Abstract
Spatial positions of cells in tissues strongly influence function, yet a high-throughput, genome-wide readout of gene expression with cellular resolution is lacking. We developed Slide-seq, a method for transferring RNA from tissue sections onto a surface covered in DNA-barcoded beads with known positions, allowing the locations of the RNA to be inferred by sequencing. Using Slide-seq, we localized cell types identified by single-cell RNA sequencing datasets within the cerebellum and hippocampus, characterized spatial gene expression patterns in the Purkinje layer of mouse cerebellum, and defined the temporal evolution of cell type-specific responses in a mouse model of traumatic brain injury. These studies highlight how Slide-seq provides a scalable method for obtaining spatially resolved gene expression data at resolutions comparable to the sizes of individual cells.
Copyright © 2019 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Figures

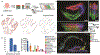


Comment in
-
Spatial transcriptomics coming of age.Nat Rev Genet. 2019 Jun;20(6):317. doi: 10.1038/s41576-019-0129-z. Nat Rev Genet. 2019. PMID: 30980030 No abstract available.
-
Pinpointing a spatial address for RNA profiles in tissues.Nature. 2019 May;569(7755):197-199. doi: 10.1038/d41586-019-01405-1. Nature. 2019. PMID: 31053836 No abstract available.
-
Slide-Seq for Spatially Mapping Gene Expression. Metabolic Syndrome Exacerbates Group 2 Pulmonary Hypertension, and NAD Metabolism Is Influenced by Tissue Origin.Am J Respir Cell Mol Biol. 2020 Jan;62(1):112-114. doi: 10.1165/rcmb.2019-0333RO. Am J Respir Cell Mol Biol. 2020. PMID: 31633380 No abstract available.
References
References and Notes
-
- Shah S, Lubeck E, Zhou W, Cai L, seqFISH Accurately Detects Transcripts in Single Cells and Reveals Robust Spatial Organization in the Hippocampus. Neuron. 94, 752–758.e1 (2017). - PubMed
-
- Stahl PL et al., Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science. 353, 78–82 (2016). - PubMed
Supplemental References:
-
- Robinson GA, Immediate early gene expression in axotomized and regenerating retinal ganglion cells of the adult rat. Mol. Brain Res. 24, 43–54 (1994). - PubMed
-
- Honkaniemi J, Sagar SM, Pyykönen I, Hicks KJ, Sharp FR, Focal brain injury induces multiple immediate early genes encoding zinc finger transcription factors. Mol. Brain Res. 28, 157–163 (1995). - PubMed
-
- Lowe DG, in Proceedings of the Seventh IEEE International Conference on Computer Vision (IEEE, 1999; http://ieeexplore.ieee.org/document/790410/), pp. 1150–1157 vol.2.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

