Dynamic changes to lipid mediators support transitions among macrophage subtypes during muscle regeneration
- PMID: 30936495
- PMCID: PMC6537107
- DOI: 10.1038/s41590-019-0356-7
Dynamic changes to lipid mediators support transitions among macrophage subtypes during muscle regeneration
Erratum in
-
Publisher Correction: Dynamic changes to lipid mediators support transitions among macrophage subtypes during muscle regeneration.Nat Immunol. 2019 Jun;20(6):765-767. doi: 10.1038/s41590-019-0401-6. Nat Immunol. 2019. PMID: 31048759
Abstract
Muscle damage elicits a sterile immune response that facilitates complete regeneration. Here, we used mass spectrometry-based lipidomics to map the mediator lipidome during the transition from inflammation to resolution and regeneration in skeletal muscle injury. We observed temporal regulation of glycerophospholipids and production of pro-inflammatory lipid mediators (for example, leukotrienes and prostaglandins) and specialized pro-resolving lipid mediators (for example, resolvins and lipoxins) that were modulated by ibuprofen. These time-dependent profiles were recapitulated in sorted neutrophils and Ly6Chi and Ly6Clo muscle-infiltrating macrophages, with a distinct pro-resolving signature observed in Ly6Clo macrophages. RNA sequencing of macrophages stimulated with resolvin D2 showed similarities to transcriptional changes found during the temporal transition from Ly6Chi macrophage to Ly6Clo macrophage. In vivo, resolvin D2 increased Ly6Clo macrophages and functional improvement of the regenerating muscle. These results reveal dynamic lipid mediator signatures of innate immune cells and provide a proof of concept for their exploitable effector roles in muscle regeneration.
Conflict of interest statement
Competing interests
None of the authors has any conflicts of interest to disclose.
Figures





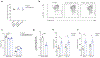
References
-
- Chazaud B Macrophages: Supportive cells for tissue repair and regeneration. Immunobiology 219, 172–178 (2014). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

