A Proteomic Screen of Neuronal Cell-Surface Molecules Reveals IgLONs as Structurally Conserved Interaction Modules at the Synapse
- PMID: 30956130
- PMCID: PMC6609445
- DOI: 10.1016/j.str.2019.03.004
A Proteomic Screen of Neuronal Cell-Surface Molecules Reveals IgLONs as Structurally Conserved Interaction Modules at the Synapse
Abstract
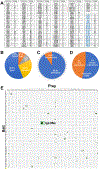
In the developing brain, cell-surface proteins play crucial roles, but their protein-protein interaction network remains largely unknown. A proteomic screen identified 200 interactions, 89 of which were not previously published. Among these interactions, we find that the IgLONs, a family of five cell-surface neuronal proteins implicated in various human disorders, interact as homo- and heterodimers. We reveal their interaction patterns and report the dimeric crystal structures of Neurotrimin (NTRI), IgLON5, and the neuronal growth regulator 1 (NEGR1)/IgLON5 complex. We show that IgLONs maintain an extended conformation and that their dimerization occurs through the first Ig domain of each monomer and is Ca2+ independent. Cell aggregation shows that NTRI and NEGR1 homo- and heterodimerize in trans. Taken together, we report 89 unpublished cell-surface ligand-receptor pairs and describe structural models of trans interactions of IgLONs, showing that their structures are compatible with a model of interaction across the synaptic cleft.
Keywords: ELISA; IgLON; SAXS; ligand-receptor pair; protein crystallography.
Copyright © 2019 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of Interests
G.T.M. is a founder of Nexomics Biosciences, Inc.
Figures



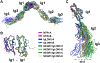


Comment in
-
Adhesion, Catalysis and Signaling: A Commonality of Association Followed by Distinctive Events Driving Function.Structure. 2019 Jul 2;27(7):1055-1056. doi: 10.1016/j.str.2019.06.003. Structure. 2019. PMID: 31269459
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

