CRISPR-Cas ribonucleoprotein mediated homology-directed repair for efficient targeted genome editing in microalgae Nannochloropsis oceanica IMET1
- PMID: 30962821
- PMCID: PMC6432748
- DOI: 10.1186/s13068-019-1401-3
CRISPR-Cas ribonucleoprotein mediated homology-directed repair for efficient targeted genome editing in microalgae Nannochloropsis oceanica IMET1
Abstract
Background: Microalgae are considered as a sustainable feedstock for the production of biofuels and other value-added compounds. In particular, Nannochloropsis spp. stand out from other microalgal species due to their capabilities to accumulate both triacylglycerol (TAG) and polyunsaturated fatty acids (PUFAs). However, the commercialization of microalgae-derived products is primarily hindered by the high production costs compared to less sustainable alternatives. Efficient genome editing techniques leading to effective metabolic engineering could result in strains with enhanced productivities of interesting metabolites and thereby reduce the production costs. Competent CRISPR-based genome editing techniques have been reported in several microalgal species, and only very recently in Nannochloropsis spp. (2017). All the reported CRISPR-Cas-based systems in Nannochloropsis spp. rely on plasmid-borne constitutive expression of Cas9 and a specific guide, combined with repair of double-stranded breaks (DSB) by non-homologous end joining (NHEJ) for the target gene knockout.
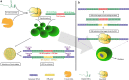
Results: In this study, we report for the first time an alternative approach for CRISPR-Cas-mediated genome editing in Nannochloropsis sp.; the Cas ribonucleoproteins (RNP) and an editing template were directly delivered into microalgal cells via electroporation, making Cas expression dispensable and homology-directed repair (HDR) possible with high efficiency. Apart from widely used SpCas9, Cas12a variants from three different bacterium were used for this approach. We observed that FnCas12a from Francisella novicida generated HDR-based targeted mutants with highest efficiency (up to 93% mutants among transformants) while AsCas12a from Acidaminococcus sp. resulted in the lowest efficiency. We initially show that the native homologous recombination (HR) system in N. oceanica IMET1 is not efficient for easy isolation of targeted mutants by HR. Cas9/sgRNA RNP delivery greatly enhanced HR at the target site, generating around 70% of positive mutant lines.
Conclusion: We show that the delivery of Cas RNP by electroporation can be an alternative approach to the presently reported plasmid-based Cas9 method for generating mutants of N. oceanica. The co-delivery of Cas-RNPs along with a dsDNA repair template efficiently enhanced HR at the target site, resulting in a remarkable higher percentage of positive mutant lines. Therefore, this approach can be used for efficient generation of targeted mutants in Nannochloropsis sp. In addition, we here report the activity of several Cas12a homologs in N. oceanica IMET1, identifying FnCas12a as the best performer for high efficiency targeted genome editing.
Keywords: CRISPR; Cas12a; Cas9; Genome editing; Homologous recombination; Homology-directed repair; Microalgae; Nannochloropsis; Ribonucleoproteins.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

References
-
- Sharma K, Schenk PM. Rapid induction of omega-3 fatty acids (EPA) in Nannochloropsis sp. by UV-C radiation. Biotechnol Bioeng. 2015;112(6):1243–1249. - PubMed
-
- Doan TTY, Sivaloganathan B, Obbard JP. Screening of marine microalgae for biodiesel feedstock. Biomass Bioenerg. 2011;35(7):2534–2544.
-
- Chisti Y. Biodiesel from microalgae. Biotechnol Adv. 2007;25(3):294–306. - PubMed
-
- Wen Z-Y, Chen F. Heterotrophic production of eicosapentaenoic acid by microalgae. Biotechnol Adv. 2003;21(4):273–294. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

