Immunometabolic Signatures Predict Risk of Progression to Active Tuberculosis and Disease Outcome
- PMID: 30967866
- PMCID: PMC6440524
- DOI: 10.3389/fimmu.2019.00527
Immunometabolic Signatures Predict Risk of Progression to Active Tuberculosis and Disease Outcome
Abstract
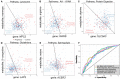
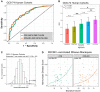
There remains a pressing need for biomarkers that can predict who will progress to active tuberculosis (TB) after exposure to Mycobacterium tuberculosis (MTB) bacterium. By analyzing cohorts of household contacts of TB index cases (HHCs) and a stringent non-human primate (NHP) challenge model, we evaluated whether integration of blood transcriptional profiling with serum metabolomic profiling can provide new understanding of disease processes and enable improved prediction of TB progression. Compared to either alone, the combined application of pre-existing transcriptome- and metabolome-based signatures more accurately predicted TB progression in the HHC cohorts and more accurately predicted disease severity in the NHPs. Pathway and data-driven correlation analyses of the integrated transcriptional and metabolomic datasets further identified novel immunometabolomic signatures significantly associated with TB progression in HHCs and NHPs, implicating cortisol, tryptophan, glutathione, and tRNA acylation networks. These results demonstrate the power of multi-omics analysis to provide new insights into complex disease processes.
Keywords: biomarker; host-pathogen interaction; household contact; inflammation; metabolomics; rhesus macaque; transcriptomics; tuberculosis.
Figures


References
-
- World Health Organization Global Tuberculosis Report. Geneva (2018). Available online at: http://www.who.int/tb/publications/global_report/en/
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

