Novel AU-rich proximal UTR sequences (APS) enhance CXCL8 synthesis upon the induction of rpS6 phosphorylation
- PMID: 30969964
- PMCID: PMC6476525
- DOI: 10.1371/journal.pgen.1008077
Novel AU-rich proximal UTR sequences (APS) enhance CXCL8 synthesis upon the induction of rpS6 phosphorylation
Abstract
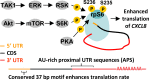
The role of ribosomal protein S6 (rpS6) phosphorylation in mRNA translation remains poorly understood. Here, we reveal a potential role in modulating the translation rate of chemokine (C-X-C motif) ligand 8 (CXCL8 or Interleukin 8, IL8). We observed that more CXCL8 protein was being secreted from less CXCL8 mRNA in primary macrophages and macrophage-like HL-60 cells relative to other cell types. This correlated with an increase in CXCL8 polyribosome association, suggesting an increase in the rate of CXCL8 translation in macrophages. The cell type-specific expression levels were replicated by a CXCL8- UTR-reporter (Nanoluc reporter flanked by the 5' and 3' UTR of CXCL8). Mutations of the CXCL8-UTR-reporter revealed that cell type-specific expression required: 1) a 3' UTR of at least three hundred bases; and 2) an AU base content that exceeds fifty percent in the first hundred bases of the 3' UTR immediately after the stop codon, which we dub AU-rich proximal UTR sequences (APS). The 5' UTR of CXCL8 enhanced expression at the protein level and conferred cell type-specific expression when paired with a 3' UTR. A search for other APS-positive mRNAs uncovered TNF alpha induced protein 6 (TNFAIP6), another mRNA that was translationally upregulated in macrophages. The elevated translation of APS-positive mRNAs in macrophages coincided with elevated rpS6 S235/236 phosphorylation. Both were attenuated by the ERK1/2 signaling inhibitors, U0126 and AZD6244. In A549 cells, rpS6 S235/236 phosphorylation was induced by TAK1, Akt or PKA signaling. This enhanced the translation of the CXCL8-UTR-reporters. Thus, we propose that the induction of rpS6 S235/236 phosphorylation enhances the translation of mRNAs that contain APS motifs, such as CXCL8 and TNFAIP6. This may contribute to the role of macrophages as the primary producer of CXCL8, a cytokine that is essential for immune cell recruitment and activation.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures









Similar articles
-
Phosphorylation of ribosomal protein S6 differentially affects mRNA translation based on ORF length.Nucleic Acids Res. 2021 Dec 16;49(22):13062-13074. doi: 10.1093/nar/gkab1157. Nucleic Acids Res. 2021. PMID: 34871442 Free PMC article.
-
Amino acid-induced translation of TOP mRNAs is fully dependent on phosphatidylinositol 3-kinase-mediated signaling, is partially inhibited by rapamycin, and is independent of S6K1 and rpS6 phosphorylation.Mol Cell Biol. 2001 Dec;21(24):8671-83. doi: 10.1128/MCB.21.24.8671-8683.2001. Mol Cell Biol. 2001. PMID: 11713299 Free PMC article.
-
Transcriptional and posttranscriptional regulation of CXCL8/IL-8 gene expression induced by connective tissue growth factor.Immunol Res. 2016 Apr;64(2):369-84. doi: 10.1007/s12026-015-8670-0. Immunol Res. 2016. PMID: 26071024
-
Ribosomal protein S6 phosphorylation: from protein synthesis to cell size.Trends Biochem Sci. 2006 Jun;31(6):342-8. doi: 10.1016/j.tibs.2006.04.003. Epub 2006 May 6. Trends Biochem Sci. 2006. PMID: 16679021 Review.
-
Ribosomal Protein S6 Phosphorylation: Four Decades of Research.Int Rev Cell Mol Biol. 2015;320:41-73. doi: 10.1016/bs.ircmb.2015.07.006. Epub 2015 Aug 5. Int Rev Cell Mol Biol. 2015. PMID: 26614871 Review.
Cited by
-
Inflammation suppresses DLG2 expression decreasing inflammasome formation.J Cancer Res Clin Oncol. 2022 Sep;148(9):2295-2311. doi: 10.1007/s00432-022-04029-7. Epub 2022 May 2. J Cancer Res Clin Oncol. 2022. PMID: 35499706 Free PMC article. Clinical Trial.
-
Alternative splicing of its 5'-UTR limits CD20 mRNA translation and enables resistance to CD20-directed immunotherapies.Blood. 2023 Nov 16;142(20):1724-1739. doi: 10.1182/blood.2023020400. Blood. 2023. PMID: 37683180 Free PMC article.
-
Role of chemokines in the crosstalk between tumor and tumor-associated macrophages.Clin Exp Med. 2023 Sep;23(5):1359-1373. doi: 10.1007/s10238-022-00888-z. Epub 2022 Sep 29. Clin Exp Med. 2023. PMID: 36173487 Free PMC article. Review.
-
Potential Therapeutic Mechanism of Scutellaria baicalensis Georgi against Ankylosing Spondylitis Based on a Comprehensive Pharmacological Model.Biomed Res Int. 2022 Dec 21;2022:9887012. doi: 10.1155/2022/9887012. eCollection 2022. Biomed Res Int. 2022. PMID: 36588535 Free PMC article.
-
[The role of IL-8 in the chronic airway inflammation and its research progress].Lin Chuang Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 2021 Dec;35(12):1144-1148. doi: 10.13201/j.issn.2096-7993.2021.12.020. Lin Chuang Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 2021. PMID: 34886635 Free PMC article. Review. Chinese.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

