The natural history of Get3-like chaperones
- PMID: 30972921
- PMCID: PMC6593721
- DOI: 10.1111/tra.12643
The natural history of Get3-like chaperones
Abstract
Get3 in yeast or TRC40 in mammals is an ATPase that, in eukaryotes, is a central element of the GET or TRC pathway involved in the targeting of tail-anchored proteins. Get3 has also been shown to possess chaperone holdase activity. A bioinformatic assessment was performed across all domains of life on functionally important regions of Get3 including the TRC40-insert and the hydrophobic groove essential for tail-anchored protein binding. We find that such a hydrophobic groove is much more common in bacterial Get3 homologs than previously appreciated based on a directed comparison of bacterial ArsA and yeast Get3. Furthermore, our analysis shows that the region containing the TRC40-insert varies in length and methionine content to an unexpected extent within eukaryotes and also between different phylogenetic groups. In fact, since the TRC40-insert is present in all domains of life, we suggest that its presence does not automatically predict a tail-anchored protein targeting function. This opens up a new perspective on the function of organellar Get3 homologs in plants which feature the TRC40-insert but have not been demonstrated to function in tail-anchored protein targeting. Our analysis also highlights a large diversity of the ways Get3 homologs dimerize. Thus, based on the structural features of Get3 homologs, these proteins may have an unexplored functional diversity in all domains of life.
Keywords: Chlorophyta; Embryophyta; Get3p; Rhodophyta; bacteria; endoplasmic reticulum; molecular chaperone; tail-anchored protein.
© 2019 The Authors. Traffic published by John Wiley & Sons Ltd.
Figures

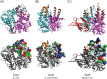



References
-
- Kutay U, Hartmann E, Rapoport TA. A class of membrane proteins with a C‐terminal anchor. Trends Cell Biol. 1993;3:72‐75. - PubMed
-
- Borgese N, Fasana E. Targeting pathways of C‐tail‐anchored proteins. Biochim Biophys Acta Biomembr. 2011;1808(3):937‐946. - PubMed
-
- Elgersma Y, Kwast L, van den Berg M, et al. Overexpression of Pex15p, a phosphorylated peroxisomal integral membrane protein required for peroxisome assembly in S.Cerevisiae, causes proliferation of the endoplasmic reticulum membrane. EMBO J. 1997;16(24):7326‐7341. 10.1093/emboj/16.24.7326. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

