Experimental Validation of the ALLNOX Program for Studying Protein-Nucleic Acid Complexes
- PMID: 30978022
- PMCID: PMC6486822
- DOI: 10.1021/acs.jpca.9b01027
Experimental Validation of the ALLNOX Program for Studying Protein-Nucleic Acid Complexes
Abstract
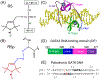
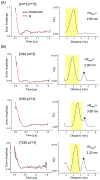
Measurement of distances between spectroscopic labels (e.g., spin labels, fluorophores) attached to specific sites of biomolecules is an important method for studying biomolecular complexes. ALLNOX (Addition of Labels and Linkers) has been developed as a program to model interlabel distances based on an input macromolecule structure. Here, we report validation of ALLNOX using measured distances between nitroxide spin labels attached to specific sites of a protein-DNA complex. The results demonstrate that ALLNOX predicts average interspin distances that matched with values measured with pairs of labels attached at the protein and/or DNA. This establishes a solid foundation for using spin labeling in conjunction with ALLNOX to investigate complexes without high-resolution structures. With its high degree of flexibility for the label or the target biomolecule, ALLNOX provides a useful tool for investigating the structure-function relationship in a large variety of biological molecules.
Figures

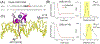
Similar articles
-
Computer Modeling of Spin Labels: NASNOX, PRONOX, and ALLNOX.Methods Enzymol. 2015;563:569-93. doi: 10.1016/bs.mie.2015.07.021. Epub 2015 Aug 31. Methods Enzymol. 2015. PMID: 26478499
-
Determination of interspin distances between spin labels attached to insulin: comparison of electron paramagnetic resonance data with the X-ray structure.Biophys J. 1997 Dec;73(6):3287-98. doi: 10.1016/S0006-3495(97)78353-X. Biophys J. 1997. PMID: 9414239 Free PMC article.
-
An Integrated Spin-Labeling/Computational-Modeling Approach for Mapping Global Structures of Nucleic Acids.Methods Enzymol. 2015;564:427-53. doi: 10.1016/bs.mie.2015.07.007. Epub 2015 Aug 8. Methods Enzymol. 2015. PMID: 26477260 Free PMC article.
-
Studying biomolecular complexes with pulsed electron-electron double resonance spectroscopy.Biochem Soc Trans. 2011 Jan;39(1):128-39. doi: 10.1042/BST0390128. Biochem Soc Trans. 2011. PMID: 21265760 Review.
-
Site-directed spin labeling studies on nucleic acid structure and dynamics.Prog Nucleic Acid Res Mol Biol. 2008;82:147-97. doi: 10.1016/S0079-6603(08)00005-6. Prog Nucleic Acid Res Mol Biol. 2008. PMID: 18929141 Free PMC article. Review.
Cited by
-
EPR Spectroscopy Provides New Insights into Complex Biological Reaction Mechanisms.J Phys Chem B. 2022 Oct 6;126(39):7486-7494. doi: 10.1021/acs.jpcb.2c05235. Epub 2022 Sep 22. J Phys Chem B. 2022. PMID: 36137278 Free PMC article. Review.
-
Site-Specific Labeling Reveals Cas9 Induces Partial Unwinding Without RNA/DNA Pairing in Sequences Distal to the PAM.CRISPR J. 2022 Apr;5(2):341-352. doi: 10.1089/crispr.2021.0100. Epub 2022 Mar 23. CRISPR J. 2022. PMID: 35352981 Free PMC article.
-
2'-Alkynyl spin-labelling is a minimally perturbing tool for DNA structural analysis.Nucleic Acids Res. 2020 Apr 6;48(6):2830-2840. doi: 10.1093/nar/gkaa086. Nucleic Acids Res. 2020. PMID: 32052020 Free PMC article.
-
A DNA unwinding equilibrium serves as a checkpoint for CRISPR-Cas12a target discrimination.Nucleic Acids Res. 2023 Sep 8;51(16):8730-8743. doi: 10.1093/nar/gkad636. Nucleic Acids Res. 2023. PMID: 37522352 Free PMC article.
-
Design of the elusive proteinaceous oxygen donor copper site suggests a promising future for copper for MRI contrast agents.Proc Natl Acad Sci U S A. 2023 Jul 4;120(27):e2219036120. doi: 10.1073/pnas.2219036120. Epub 2023 Jun 26. Proc Natl Acad Sci U S A. 2023. PMID: 37364102 Free PMC article.
References
-
- Hubbell WL; Altenbach C Investigation of structure and dynamics in membrane proteins using site-directed spin labeling. Curr. Opin. Struct. Biol 1994, 4, 566–573.
-
- Ding Y; Nguyen P; Tangprasertchai NS; Reyes CV; Zhang X; Qin PZ Nucleic acid structure and dynamics: perspectives from site-directed spin labeling In Electron Paramagnetic Resonance; The Royal Society of Chemistry, 2015; Vol. 24, pp 122–147.
-
- Bagryanskaya EG; Krumkacheva OA; Fedin MV; Marque SRA Development and application of spin traps, spin probes, and spin labels In Methods Enzymol; Qin PZ, Warncke K, Eds.; Academic Press, 2015; Vol. 563, pp 365–396. - PubMed
-
- Feintuch A; Otting G; Goldfarb D Gd3+ spin labeling for measuring distances in biomacromolecules: why and how? In Methods Enzymol; Qin PZ, Warncke K, Eds.; Academic Press, 2015; Vol. 563, pp 415–457. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

