Two new structural mutations in the 5' region of the ASIP gene cause diluted feather color phenotypes in Japanese quail
- PMID: 30987584
- PMCID: PMC6466734
- DOI: 10.1186/s12711-019-0458-6
Two new structural mutations in the 5' region of the ASIP gene cause diluted feather color phenotypes in Japanese quail
Abstract
Background: In quail, two feather colour phenotypes i.e. fawn-2/beige and yellow are associated with the ASIP locus. The aim of our study was to characterize the structural modifications within this locus that explain the yellow mutation (large deletion) and the fawn-2/beige mutation (assumed to be caused by a different structural modification).
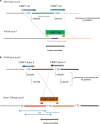
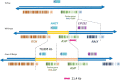
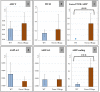
Results: For the yellow phenotype, we identified a complex mutation that involves a 141,162-bp long deletion. For the fawn-2/beige phenotype, we identified a 71-kb tandem duplication that comprises one unchanged copy of ASIP and one copy present in the ITCH-ASIP fusion gene, which leads to a transcript coding for a normal ASIP protein. Although this agrees with previous reports that reported an increased level of ASIP transcripts in the skin of mutant animals, we show that in the skin from fawn-2/beige embryos, this level is higher than expected with a simple duplication of the ASIP gene. Thus, we hypothesize that the 5' region of the ITCH-ASIP fusion gene leads to a higher transcription level than the 5' region of the ASIP gene.
Conclusions: We were able to conclude that the fawn-2 and beige phenotypes are caused by the same allele at the ASIP locus. Both of the associated mutations fawn-2/beige and yellow lead to the formation of a fusion gene, which encodes a transcript for the ASIP protein. In both cases, transcription of ASIP depends on the promoter of a different gene, which includes alternative up-regulating sequences. However, we cannot exclude the possibility that the loss of the 5' region of the ASIP gene itself has additional impacts, especially for the fawn-2/beige mutation. In addition, in several other species including mammals, the existence of other dominant gain-of-function structural modifications that are localized upstream of the ASIP coding sequences has been reported, which supports our hypothesis that repressors in the 5' region of ASIP are absent in the fawn-2/beige mutant.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Characterization of Japanese quail yellow as a genomic deletion upstream of the avian homolog of the mammalian ASIP (agouti) gene.Genetics. 2008 Feb;178(2):777-86. doi: 10.1534/genetics.107.077073. Genetics. 2008. PMID: 18287407 Free PMC article.
-
Recessive black is allelic to the yellow plumage locus in Japanese quail and associated with a frameshift deletion in the ASIP gene.Genetics. 2008 Feb;178(2):771-5. doi: 10.1534/genetics.107.077040. Genetics. 2008. PMID: 18287406 Free PMC article.
-
A gene duplication affecting expression of the ovine ASIP gene is responsible for white and black sheep.Genome Res. 2008 Aug;18(8):1282-93. doi: 10.1101/gr.072090.107. Epub 2008 May 20. Genome Res. 2008. PMID: 18493018 Free PMC article.
-
Evolutionary and phylogeographic views on Mc1r and Asip variation in mammals.Genes Genet Syst. 2013;88(3):155-64. doi: 10.1266/ggs.88.155. Genes Genet Syst. 2013. PMID: 24025244 Review.
-
The untranslated side of hair and skin mammalian pigmentation: Beyond coding sequences.IUBMB Life. 2010 May;62(5):340-6. doi: 10.1002/iub.318. IUBMB Life. 2010. PMID: 20222017 Review.
Cited by
-
A 13.42-kb tandem duplication at the ASIP locus is strongly associated with the depigmentation phenotype of non-classic Swiss markings in goats.BMC Genomics. 2022 Jun 13;23(1):437. doi: 10.1186/s12864-022-08672-9. BMC Genomics. 2022. PMID: 35698044 Free PMC article.
-
A high-quality assembly reveals genomic characteristics, phylogenetic status, and causal genes for leucism plumage of Indian peafowl.Gigascience. 2022 Apr 6;11:giac018. doi: 10.1093/gigascience/giac018. Gigascience. 2022. PMID: 35383847 Free PMC article.
-
Understanding the skin blackening phenomenon in Youzhou Dark goats based on the histological characteristics of melanocytes.J Adv Vet Anim Res. 2024 Dec 29;11(4):1139-1148. doi: 10.5455/javar.2024.k865. eCollection 2024 Dec. J Adv Vet Anim Res. 2024. PMID: 40013286 Free PMC article.
-
Genetic Basis and Evolution of Structural Color Polymorphism in an Australian Songbird.Mol Biol Evol. 2024 Mar 1;41(3):msae046. doi: 10.1093/molbev/msae046. Mol Biol Evol. 2024. PMID: 38415852 Free PMC article.
-
Whole-genome selection signature differences between Chaohu and Ji'an red ducks.BMC Genomics. 2024 May 27;25(1):522. doi: 10.1186/s12864-024-10339-6. BMC Genomics. 2024. PMID: 38802792 Free PMC article.
References
-
- Fontanesi L, Forestier L, Allain D, Scotti E, Beretti F, Deretz-Picoulet S, et al. Characterization of the rabbit agouti signaling protein (ASIP) gene: transcripts and phylogenetic analyses and identification of the causative mutation of the nonagouti black coat colour. Genomics. 2010;95:166–175. doi: 10.1016/j.ygeno.2009.11.003. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

