Proteasome dysfunction induces excessive proteome instability and loss of mitostasis that can be mitigated by enhancing mitochondrial fusion or autophagy
- PMID: 31002009
- PMCID: PMC6735541
- DOI: 10.1080/15548627.2019.1596477
Proteasome dysfunction induces excessive proteome instability and loss of mitostasis that can be mitigated by enhancing mitochondrial fusion or autophagy
Abstract
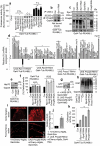
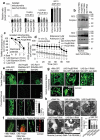
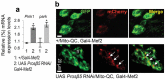
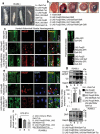
The ubiquitin-proteasome pathway (UPP) is central to proteostasis network (PN) functionality and proteome quality control. Yet, the functional implication of the UPP in tissue homeodynamics at the whole organism level and its potential cross-talk with other proteostatic or mitostatic modules are not well understood. We show here that knock down (KD) of proteasome subunits in Drosophila flies, induced, for most subunits, developmental lethality. Ubiquitous or tissue specific proteasome dysfunction triggered systemic proteome instability and activation of PN modules, including macroautophagy/autophagy, molecular chaperones and the antioxidant cncC (the fly ortholog of NFE2L2/Nrf2) pathway. Also, proteasome KD increased genomic instability, altered metabolic pathways and severely disrupted mitochondrial functionality, triggering a cncC-dependent upregulation of mitostatic genes and enhanced rates of mitophagy. Whereas, overexpression of key regulators of antioxidant responses (e.g., cncC or foxo) could not suppress the deleterious effects of proteasome dysfunction; these were alleviated in both larvae and adult flies by modulating mitochondrial dynamics towards increased fusion or by enhancing autophagy. Our findings reveal the extensive functional wiring of genomic, proteostatic and mitostatic modules in higher metazoans. Also, they support the notion that age-related increase of proteotoxic stress due to decreased UPP activity deregulates all aspects of cellular functionality being thus a driving force for most age-related diseases. Abbreviations: ALP: autophagy-lysosome pathway; ARE: antioxidant response element; Atg8a: autophagy-related 8a; ATPsynβ: ATP synthase, β subunit; C-L: caspase-like proteasomal activity; cncC: cap-n-collar isoform-C; CT-L: chymotrypsin-like proteasomal activity; Drp1: dynamin related protein 1; ER: endoplasmic reticulum; foxo: forkhead box, sub-group O; GLU: glucose; GFP: green fluorescent protein; GLY: glycogen; Hsf: heat shock factor; Hsp: Heat shock protein; Keap1: kelch-like ECH-associated protein 1; Marf: mitochondrial assembly regulatory factor; NFE2L2/Nrf2: nuclear factor, erythroid 2 like 2; Opa1: optic atrophy 1; PN: proteostasis network; RNAi: RNA interference; ROS: reactive oxygen species; ref(2)P: refractory to sigma P; SQSTM1: sequestosome 1; SdhA: succinate dehydrogenase, subunit A; T-L: trypsin-like proteasomal activity; TREH: trehalose; UAS: upstream activation sequence; Ub: ubiquitin; UPR: unfolded protein response; UPP: ubiquitin-proteasome pathway.
Keywords: Aging; autophagy; cncC; foxo; mitostasis; proteasome; proteostasis.
Figures
Similar articles
-
Functional wiring of proteostatic and mitostatic modules ensures transient organismal survival during imbalanced mitochondrial dynamics.Redox Biol. 2019 Jun;24:101219. doi: 10.1016/j.redox.2019.101219. Epub 2019 May 17. Redox Biol. 2019. PMID: 31132524 Free PMC article.
-
Mitochondrial protein import regulates cytosolic protein homeostasis and neuronal integrity.Autophagy. 2018;14(8):1293-1309. doi: 10.1080/15548627.2018.1474991. Epub 2018 Jul 21. Autophagy. 2018. PMID: 29909722 Free PMC article.
-
Limiting cap-dependent translation increases 20S proteasomal degradation and protects the proteomic integrity in autophagy-deficient skeletal muscle.Autophagy. 2025 Jun;21(6):1212-1227. doi: 10.1080/15548627.2025.2457925. Epub 2025 Feb 6. Autophagy. 2025. PMID: 39878121 Free PMC article.
-
Molecular chaperones and proteostasis regulation during redox imbalance.Redox Biol. 2014 Jan 30;2:323-32. doi: 10.1016/j.redox.2014.01.017. eCollection 2014. Redox Biol. 2014. PMID: 24563850 Free PMC article. Review.
-
The amazing ubiquitin-proteasome system: structural components and implication in aging.Int Rev Cell Mol Biol. 2015;314:171-237. doi: 10.1016/bs.ircmb.2014.09.002. Epub 2014 Nov 18. Int Rev Cell Mol Biol. 2015. PMID: 25619718 Review.
Cited by
-
Cardiovascular Toxicity of Proteasome Inhibitors: Underlying Mechanisms and Management Strategies: JACC: CardioOncology State-of-the-Art Review.JACC CardioOncol. 2023 Feb 21;5(1):1-21. doi: 10.1016/j.jaccao.2022.12.005. eCollection 2023 Feb. JACC CardioOncol. 2023. PMID: 36875897 Free PMC article. Review.
-
NFE2L1 as a central regulator of proteostasis in neurodegenerative diseases: interplay with autophagy, ferroptosis, and the proteasome.Front Mol Neurosci. 2025 May 1;18:1551571. doi: 10.3389/fnmol.2025.1551571. eCollection 2025. Front Mol Neurosci. 2025. PMID: 40375958 Free PMC article. Review.
-
Clusterin overexpression in mice exacerbates diabetic phenotypes but suppresses tumor progression in a mouse melanoma model.Aging (Albany NY). 2021 Mar 10;13(5):6485-6505. doi: 10.18632/aging.202788. Epub 2021 Mar 10. Aging (Albany NY). 2021. PMID: 33744871 Free PMC article.
-
Nrf2 activation induces mitophagy and reverses Parkin/Pink1 knock down-mediated neuronal and muscle degeneration phenotypes.Cell Death Dis. 2021 Jul 3;12(7):671. doi: 10.1038/s41419-021-03952-w. Cell Death Dis. 2021. PMID: 34218254 Free PMC article.
-
Boosting autophagy in anti-tumor proteasome inhibition-mediated cardiotoxicity.Aging (Albany NY). 2023 May 8;15(9):3226-3227. doi: 10.18632/aging.204724. Epub 2023 May 8. Aging (Albany NY). 2023. PMID: 37166427 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
