Growth Hormone Induces Colon DNA Damage Independent of IGF-1
- PMID: 31002310
- PMCID: PMC6530523
- DOI: 10.1210/en.2019-00132
Growth Hormone Induces Colon DNA Damage Independent of IGF-1
Abstract
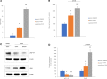
DNA damage occurs as a result of environmental insults and aging and, if unrepaired, may lead to chromosomal instability and tumorigenesis. Because GH suppresses ataxia-telangiectasia mutated kinase phosphorylation, decreases DNA repair, and increases DNA damage accumulation, we elucidated whether GH effects on DNA damage are mediated through induced IGF-1. In nontumorous human colon cells, GH, but not IGF-1, increased DNA damage. Stably disrupted IGF-1 receptor (IGF-1R) by lentivirus-expressing short hairpin RNA in vitro or treatment with the IGF-1R phosphorylation inhibitor picropodophyllotoxin (PPP) in vitro and in vivo led to markedly induced GH receptor (GHR) abundance, rendering cells more responsive to GH actions. Suppressing IGF-1R triggered DNA damage in both normal human colon cells and three-dimensional human intestinal organoids. DNA damage was further increased when cells with disrupted IGF-1R were treated with GH. Because GH induction of DNA damage accumulation appeared to be mediated not by IGF-1R but probably by more abundant GH receptor expression, we injected athymic mice with GH-secreting xenografts and then treated them with PPP. In these mice, high circulating GH levels were associated with increased colon DNA damage despite disrupted IGF-1R activity (P < 0.01), whereas GHR levels were also induced. Further confirming that GH effects on DNA damage are directly mediated by GHR signaling, GHR-/- mice injected with PPP did not show increased DNA damage, whereas wild-type mice with intact GHR exhibited increased colon DNA damage in the face of IGF-1 signaling suppression. The results indicate that GH directly induces DNA damage independent of IGF-1.
Copyright © 2019 Endocrine Society.
Figures


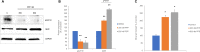


Similar articles
-
Decreases in growth hormone receptor signal transduction contribute to the decline in insulin-like growth factor I gene expression with age.Endocrinology. 1995 Oct;136(10):4551-7. doi: 10.1210/endo.136.10.7664676. Endocrinology. 1995. PMID: 7664676
-
Human GH receptor-IGF-1 receptor interaction: implications for GH signaling.Mol Endocrinol. 2014 Nov;28(11):1841-54. doi: 10.1210/me.2014-1174. Epub 2014 Sep 11. Mol Endocrinol. 2014. PMID: 25211187 Free PMC article.
-
Craniopharyngioma cell growth is promoted by growth hormone (GH) and is inhibited by tamoxifen: involvement of growth hormone receptor (GHR) and IGF-1 receptor (IGF-1R).J Clin Neurosci. 2013 Jan;20(1):153-7. doi: 10.1016/j.jocn.2012.04.014. Epub 2012 Oct 30. J Clin Neurosci. 2013. PMID: 23117141
-
Impact of growth hormone hypersecretion on the adult human kidney.Ann Endocrinol (Paris). 2011 Dec;72(6):485-95. doi: 10.1016/j.ando.2011.08.001. Epub 2011 Nov 17. Ann Endocrinol (Paris). 2011. PMID: 22098791 Review.
-
Insulin-like growth factor 1 (IGF-1) and aging: controversies and new insights.Biogerontology. 2003;4(1):1-8. doi: 10.1023/a:1022448532248. Biogerontology. 2003. PMID: 12652183 Review.
Cited by
-
Benefits of Living Without Growth Hormone.J Gerontol A Biol Sci Med Sci. 2021 Sep 13;76(10):1769-1774. doi: 10.1093/gerona/glab147. J Gerontol A Biol Sci Med Sci. 2021. PMID: 34036341 Free PMC article. No abstract available.
-
Pituitary tumors and the risk of other malignancies: is the relationship coincidental or causal?Endocr Oncol. 2022 Dec 22;2(1):R1-R13. doi: 10.1530/EO-21-0033. eCollection 2022 Jan. Endocr Oncol. 2022. PMID: 37435457 Free PMC article. Review.
-
Non-pituitary growth hormone enables colon cell senescence evasion.Aging Cell. 2024 Aug;23(8):e14193. doi: 10.1111/acel.14193. Epub 2024 May 9. Aging Cell. 2024. PMID: 38724466 Free PMC article.
-
Covert actions of growth hormone: fibrosis, cardiovascular diseases and cancer.Nat Rev Endocrinol. 2022 Sep;18(9):558-573. doi: 10.1038/s41574-022-00702-6. Epub 2022 Jun 24. Nat Rev Endocrinol. 2022. PMID: 35750929 Free PMC article. Review.
-
Peptide Hormone Regulation of DNA Damage Responses.Endocr Rev. 2020 Jul 1;41(4):bnaa009. doi: 10.1210/endrev/bnaa009. Endocr Rev. 2020. PMID: 32270196 Free PMC article. Review.
References
-
- Clayton PE, Gill MS, Tillmann V, Westwood M. Translational neuroendocrinology: control of human growth. J Neuroendocrinol. 2014;26(6):349–355. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

