Dopamine receptor D4 (DRD4) polymorphisms with reduced functional potency intensify atrophy in syndrome-specific sites of frontotemporal dementia
- PMID: 31003069
- PMCID: PMC6475809
- DOI: 10.1016/j.nicl.2019.101822
Dopamine receptor D4 (DRD4) polymorphisms with reduced functional potency intensify atrophy in syndrome-specific sites of frontotemporal dementia
Abstract
Objective: We aimed to understand the impact of dopamine receptor D4 (DRD4) polymorphisms on neurodegeneration in patients with dementia. We hypothesized that DRD4dampened-variants with reduced functional potency would be associated with greater atrophy in regions with higher receptor density. Given that DRD4 is concentrated in anterior regions of the limbic and cortical forebrain we anticipated genotype effects in patients with a more rostral pattern of neurodegeneration.
Methods: 337 subjects, including healthy controls, patients with Alzheimer's disease (AD) and frontotemporal dementia (FTD) underwent genotyping, structural MRI, and cognitive/behavioral testing. We conducted whole-brain voxel-based morphometry to examine the relationship between DRD4 genotypes and brain atrophy patterns within and across groups. General linear modeling was used to evaluate relationships between genotype and cognitive/behavioral measures.
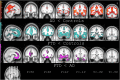
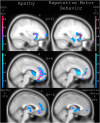
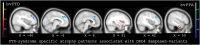
Results: DRD4 dampened-variants predicted gray matter atrophy in disease-specific regions of FTD in anterior cingulate, ventromedial prefrontal, orbitofrontal and insular cortices on the right greater than the left. Genotype predicted greater apathy and repetitive motor disturbance in patients with FTD. These results covaried with frontoinsular cortical atrophy. Peak atrophy patterned along regions of neuroanatomic vulnerability in FTD-spectrum disorders. In AD subjects and controls, genotype did not impact gray matter intensity.
Conclusions: We conclude that DRD4 polymorphisms with reduced functional potency exacerbate neuronal injury in sites of higher receptor density, which intersect with syndrome-specific regions undergoing neurodegeneration in FTD.
Keywords: Anterior cingulate cortex; Apathy; DRD(4); Frontotemporal dementia; Insula; Salience network.
Copyright © 2019 The Authors. Published by Elsevier Inc. All rights reserved.
Figures

References
-
- Agosta F., Vossel K.A., Miller B.L., Migliaccio R., Bonasera S.J., Filippi M., Boxer A.L., Karydas A., Possin K.L., Gorno-Tempini M.L. Apolipoprotein E E4 is associated with disease-specific effects on brain atrophy in Alzheimer's disease and frontotemporal dementia. Proc. Natl. Acad. Sci. U. S. A. 2009;106:2018–2022. - PMC - PubMed
-
- Armstrong M.J., Litvan I., Lang A.E., Bak T.H., Bhatia K.P., Borroni B., Boxer A.L., Dickson D.W., Grossman M., Hallett M., Josephs K., Kerterz A., Lee S.E., Miller B.L., Reich S.G., Riley D.E., Tolosa E., Tröster A.I., Vidailhet M., Weiner W.J. Criteria for the diagnosis of corticobasal degeneration. Neurology. 2013;80:496–503. - PMC - PubMed
-
- Asghari V., Schoots O., Van Kats S., Ohara K., Jovanovic V., Guan H.-C., Bunzow J.R., Petronis A., Van Tol H.H.M. Dopamine D4 receptor repeat: analysis of different native and mutant forms of the human and rat genes. Mol. Pharm. 1994;46:364–373. - PubMed
-
- Asghari V., Sanyal S., Buchwaldt S., Paterson A., Jovanovic V., Van Tol H.H.M. Modulation of intercellular cyclic AMP levels by different human dopamine receptor variants. J. Neurochem. 1995;65:1157–1165. - PubMed
-
- Butler P.M., Matthews L.J. Novelty-seeking DRD4 polymorphisms are associated with human migration distance out-of-Africa after controlling for neutral population gene structure. Am. J. Phys. Anthr. 2011;145:382–389. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- RC1 AG035610/AG/NIA NIH HHS/United States
- U24 AG021886/AG/NIA NIH HHS/United States
- R01 AG052496/AG/NIA NIH HHS/United States
- T32 AG023481/AG/NIA NIH HHS/United States
- P01 AG019724/AG/NIA NIH HHS/United States
- R01 MH097268/MH/NIMH NIH HHS/United States
- K24 DC015544/DC/NIDCD NIH HHS/United States
- R01 AG026938/AG/NIA NIH HHS/United States
- K23 AG045289/AG/NIA NIH HHS/United States
- P50 AG023501/AG/NIA NIH HHS/United States
- U01 AG052943/AG/NIA NIH HHS/United States
- P30 NS062691/NS/NINDS NIH HHS/United States
- R01 AG057204/AG/NIA NIH HHS/United States
- R01 NS100440/NS/NINDS NIH HHS/United States
- R56 NS050915/NS/NINDS NIH HHS/United States
- R01 NS050915/NS/NINDS NIH HHS/United States

