Missing the Match Might Not Cost You the Game: Primer-Template Mismatches Studied in Different Hepatitis A Virus Variants
- PMID: 31004336
- PMCID: PMC6689102
- DOI: 10.1007/s12560-019-09387-z
Missing the Match Might Not Cost You the Game: Primer-Template Mismatches Studied in Different Hepatitis A Virus Variants
Abstract
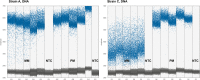
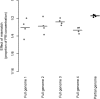
Mismatches between template sequences and reverse transcription (RT) or polymerase chain reaction (PCR) primers can lead to underestimation or false negative results during detection and quantification of sequence-diverse viruses. We performed an in silico inclusivity analysis of a widely used RT-PCR assay for detection of hepatitis A virus (HAV) in food, described in ISO 15216-1. One of the most common mismatches found was a single G (primer) to U (template) mismatch located at the terminal 3'-end of the reverse primer region. This mismatch was present in all genotype III sequences available in GenBank. Partial HAV genomes with common or potentially severe mismatches were produced by in vitro transcription and analysed using RT-ddPCR and RT-qPCR. When using standard conditions for RT-qPCR, the mismatch identified resulted in underestimation of the template concentration by a factor of 1.7-1.8 and an increase in 95% limit of detection from 8.6 to 19 copies/reaction. The effect of this mismatch was verified using full-length viral genomes. Here, the same mismatch resulted in underestimation of the template concentration by a factor of 2.8. For the partial genomes, the presence of additional mismatches resulted in underestimation of the template concentration by up to a factor of 232. Quantification by RT-ddPCR and RT-qPCR was equally affected during analysis of RNA templates with mismatches within the reverse primer region. However, on analysing DNA templates with the same mismatches, we found that ddPCR quantification was less affected by mismatches than qPCR due to the end-point detection technique.
Keywords: Digital PCR; Hepatitis A virus; Mismatch; Primer; Real-time PCR; Reverse transcription.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


Similar articles
-
Development of a One-Step Duplex RT-PCR Method for the Simultaneous Detection of VP3/VP1 and VP1/P2B Regions of the Hepatitis A Virus.J Microbiol Biotechnol. 2016 Aug 28;26(8):1398-403. doi: 10.4014/jmb.1604.04047. J Microbiol Biotechnol. 2016. PMID: 27197668
-
The effect of multiple primer-template mismatches on quantitative PCR accuracy and development of a multi-primer set assay for accurate quantification of pcrA gene sequence variants.J Microbiol Methods. 2013 Sep;94(3):224-31. doi: 10.1016/j.mimet.2013.06.013. Epub 2013 Jun 25. J Microbiol Methods. 2013. PMID: 23806694
-
The impact of primer and probe-template mismatches on the sensitivity of pandemic influenza A/H1N1/2009 virus detection by real-time RT-PCR.J Clin Virol. 2010 Jun;48(2):91-5. doi: 10.1016/j.jcv.2010.03.012. Epub 2010 Apr 21. J Clin Virol. 2010. PMID: 20413345
-
Comparison of HIV-1 and avian myeloblastosis virus reverse transcriptase fidelity on RNA and DNA templates.J Biol Chem. 1992 May 25;267(15):10888-96. J Biol Chem. 1992. PMID: 1375233
-
Brief guide to RT-qPCR.Mol Cells. 2024 Dec;47(12):100141. doi: 10.1016/j.mocell.2024.100141. Epub 2024 Oct 28. Mol Cells. 2024. PMID: 39476972 Free PMC article. Review.
Cited by
-
Hepatitis A Virus and Hepatitis E Virus as Food- and Waterborne Pathogens-Transmission Routes and Methods for Detection in Food.Viruses. 2023 Aug 12;15(8):1725. doi: 10.3390/v15081725. Viruses. 2023. PMID: 37632066 Free PMC article. Review.
-
Droplet digital PCR of viral DNA/RNA, current progress, challenges, and future perspectives.J Med Virol. 2021 Jul;93(7):4182-4197. doi: 10.1002/jmv.26846. Epub 2021 Mar 11. J Med Virol. 2021. PMID: 33538349 Free PMC article. Review.
-
Hepatitis a Vaccine as Opportunity of Primary Prevention for Food Handlers: A Narrative Review.Vaccines (Basel). 2023 Jul 21;11(7):1271. doi: 10.3390/vaccines11071271. Vaccines (Basel). 2023. PMID: 37515087 Free PMC article. Review.
-
Development of Human Rhinovirus RNA Reference Material Using Digital PCR.Genes (Basel). 2023 Dec 14;14(12):2210. doi: 10.3390/genes14122210. Genes (Basel). 2023. PMID: 38137032 Free PMC article.
-
Advances in the differential molecular diagnosis of vesicular disease pathogens in swine.Front Microbiol. 2022 Oct 25;13:1019876. doi: 10.3389/fmicb.2022.1019876. eCollection 2022. Front Microbiol. 2022. PMID: 36386633 Free PMC article. Review.
References
-
- Agresti A. Categorical data analysis. Hoboken: Wiley; 2003.
-
- Bates, D., Mächler, M., Bolker, B., & Walker, S. (2014). Fitting linear mixed-effects models using lme4. arXiv:1406.5823.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

