Effects of Noncanonical Base Pairing on RNA Folding: Structural Context and Spatial Arrangements of G·A Pairs
- PMID: 31008589
- PMCID: PMC6729125
- DOI: 10.1021/acs.biochem.9b00122
Effects of Noncanonical Base Pairing on RNA Folding: Structural Context and Spatial Arrangements of G·A Pairs
Abstract
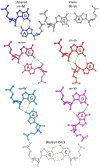
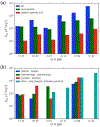
Noncanonical base pairs play important roles in assembling the three-dimensional structures critical to the diverse functions of RNA. These associations contribute to the looped segments that intersperse the canonical double-helical elements within folded, globular RNA molecules. They stitch together various structural elements, serve as recognition elements for other molecules, and act as sites of intrinsic stiffness or deformability. This work takes advantage of new software (DSSR) designed to streamline the analysis and annotation of RNA three-dimensional structures. The multiscale structural information gathered for individual molecules, combined with the growing number of unique, well-resolved RNA structures, makes it possible to examine the collective features deeply and to uncover previously unrecognized patterns of chain organization. Here we focus on a subset of noncanonical base pairs involving guanine and adenine and the links between their modes of association, secondary structural context, and contributions to tertiary folding. The rigorous descriptions of base-pair geometry that we employ facilitate characterization of recurrent geometric motifs and the structural settings in which these arrangements occur. Moreover, the numerical parameters hint at the natural motions of the interacting bases and the pathways likely to connect different spatial forms. We draw attention to higher-order multiplexes involving two or more G·A pairs and the roles these associations appear to play in bridging different secondary structural units. The collective data reveal pairing propensities in base organization, secondary structural context, and deformability and serve as a starting point for further multiscale investigations and/or simulations of RNA folding.
Figures





Similar articles
-
New information content in RNA base pairing deduced from quantitative analysis of high-resolution structures.Methods. 2009 Mar;47(3):177-86. doi: 10.1016/j.ymeth.2008.12.003. Epub 2009 Jan 14. Methods. 2009. PMID: 19150407 Free PMC article. Review.
-
The role of N7 protonation of guanine in determining the structure, stability and function of RNA base pairs.Phys Chem Chem Phys. 2015 Oct 21;17(39):26249-63. doi: 10.1039/c5cp04894j. Epub 2015 Sep 18. Phys Chem Chem Phys. 2015. PMID: 26382322
-
Structural characterization of naturally occurring RNA single mismatches.Nucleic Acids Res. 2011 Feb;39(3):1081-94. doi: 10.1093/nar/gkq793. Epub 2010 Sep 28. Nucleic Acids Res. 2011. PMID: 20876693 Free PMC article.
-
RNA structure and dynamics: a base pairing perspective.Prog Biophys Mol Biol. 2013 Nov;113(2):264-83. doi: 10.1016/j.pbiomolbio.2013.07.003. Epub 2013 Jul 23. Prog Biophys Mol Biol. 2013. PMID: 23891726 Review.
-
The G x U wobble base pair. A fundamental building block of RNA structure crucial to RNA function in diverse biological systems.EMBO Rep. 2000 Jul;1(1):18-23. doi: 10.1093/embo-reports/kvd001. EMBO Rep. 2000. PMID: 11256617 Free PMC article. Review.
Cited by
-
A tRNA-mimic Strategy to Explore the Role of G34 of tRNAGly in Translation and Codon Frameshifting.Int J Mol Sci. 2019 Aug 11;20(16):3911. doi: 10.3390/ijms20163911. Int J Mol Sci. 2019. PMID: 31405256 Free PMC article.
-
DNA as a perfect quantum computer based on the quantum physics principles.Sci Rep. 2024 May 21;14(1):11636. doi: 10.1038/s41598-024-62539-5. Sci Rep. 2024. PMID: 38773193 Free PMC article.
-
Investigating the correlation between Xrn1-resistant RNAs and frameshifter pseudoknots.RNA Biol. 2023 Jan;20(1):409-418. doi: 10.1080/15476286.2023.2205224. RNA Biol. 2023. PMID: 37400999 Free PMC article.
-
Alu RNA fold links splicing with signal recognition particle proteins.Nucleic Acids Res. 2023 Aug 25;51(15):8199-8216. doi: 10.1093/nar/gkad500. Nucleic Acids Res. 2023. PMID: 37309897 Free PMC article.
-
Antibody designing against IIIabc junction (JIIIabc) of HCV IRES through affinity maturation; RNA-Antibody docking and interaction analysis.PLoS One. 2023 Sep 8;18(9):e0291213. doi: 10.1371/journal.pone.0291213. eCollection 2023. PLoS One. 2023. PMID: 37682810 Free PMC article.
References
-
- Rich A, and RajBhandary UL (1976) Transfer RNA: molecular structure, sequence, and properties, Annu Rev Biochem 45, 805–860. - PubMed
-
- Ferré-D’Amaré AR, and Doudna JA (1999) RNA folds: insights from recent crystal structures, Annu Rev Biophys Biomol Struct 28, 57–73. - PubMed
-
- Hermann T, and Patel DJ (1999) Stitching together RNA tertiary architectures, J Mol Biol 294, 829–849. - PubMed
-
- Draper DE (1995) Protein-RNA recognition, Annu Rev Biochem 64, 593–620. - PubMed
-
- Hermann T, and Westhof E (1999) Non-Watson–Crick base pairs in RNA-protein recognition, Chem Biol 6, R335–R343. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

