Ex vivo Dynamics of Human Glioblastoma Cells in a Microvasculature-on-a-Chip System Correlates with Tumor Heterogeneity and Subtypes
- PMID: 31016107
- PMCID: PMC6468969
- DOI: 10.1002/advs.201801531
Ex vivo Dynamics of Human Glioblastoma Cells in a Microvasculature-on-a-Chip System Correlates with Tumor Heterogeneity and Subtypes
Abstract
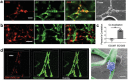
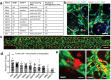
The perivascular niche (PVN) plays an essential role in brain tumor stem-like cell (BTSC) fate control, tumor invasion, and therapeutic resistance. Here, a microvasculature-on-a-chip system as a PVN model is used to evaluate the ex vivo dynamics of BTSCs from ten glioblastoma patients. BTSCs are found to preferentially localize in the perivascular zone, where they exhibit either the lowest motility, as in quiescent cells, or the highest motility, as in the invasive phenotype, with migration over long distance. These results indicate that PVN is a niche for BTSCs, while the microvascular tracks may serve as a path for tumor cell migration. The degree of colocalization between tumor cells and microvessels varies significantly across patients. To validate these results, single-cell transcriptome sequencing (10 patients and 21 750 single cells in total) is performed to identify tumor cell subtypes. The colocalization coefficient is found to positively correlate with proneural (stem-like) or mesenchymal (invasive) but not classical (proliferative) tumor cells. Furthermore, a gene signature profile including PDGFRA correlates strongly with the "homing" of tumor cells to the PVN. These findings demonstrate that the model can recapitulate in vivo tumor cell dynamics and heterogeneity, representing a new route to study patient-specific tumor cell functions.
Keywords: brain tumor dynamics; ex vivo assays; microvasculature; organ‐on‐a‐chip.
Conflict of interest statement
Rong Fan is on the Scientific Advisory Boards of IsoPlexis, Bio‐Techne, and Singleron Biotechnologies with financial interest.
Figures



References
-
- Gilbertson R. J., Rich J. N., Nat. Rev. Cancer 2007, 7, 733. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
