Agar plate-based screening methods for the identification of polyester hydrolysis by Pseudomonas species
- PMID: 31016871
- PMCID: PMC6922526
- DOI: 10.1111/1751-7915.13418
Agar plate-based screening methods for the identification of polyester hydrolysis by Pseudomonas species
Abstract
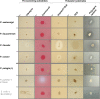
Hydrolases acting on polyesters like cutin, polycaprolactone or polyethylene terephthalate (PET) are of interest for several biotechnological applications like waste treatment, biocatalysis and sustainable polymer modifications. Recent studies suggest that a large variety of such enzymes are still to be identified and explored in a variety of microorganisms, including bacteria of the genus Pseudomonas. For activity-based screening, methods have been established using agar plates which contain nanoparticles of polycaprolactone or PET prepared by solvent precipitation and evaporation. In this protocol article, we describe a straightforward agar plate-based method using emulsifiable artificial polyesters as substrates, namely Impranil® DLN and liquid polycaprolactone diol (PLD). Thereby, the currently quite narrow set of screening substrates is expanded. We also suggest optional pre-screening with short-chain and middle-chain-length triglycerides as substrates to identify enzymes with lipolytic activity to be further tested for polyesterase activity. We applied these assays to experimentally demonstrate polyesterase activity in bacteria from the P. pertucinogena lineage originating from contaminated soils and diverse marine habitats.
© 2019 The Authors. Microbial Biotechnology published by John Wiley & Sons Ltd and Society for Applied Microbiology.
Conflict of interest statement
None declared.
Figures

Similar articles
-
Identification of BgP, a Cutinase-Like Polyesterase From a Deep-Sea Sponge-Derived Actinobacterium.Front Microbiol. 2022 Apr 12;13:888343. doi: 10.3389/fmicb.2022.888343. eCollection 2022. Front Microbiol. 2022. PMID: 35495686 Free PMC article.
-
Antarctic Polyester Hydrolases Degrade Aliphatic and Aromatic Polyesters at Moderate Temperatures.Appl Environ Microbiol. 2022 Jan 11;88(1):e0184221. doi: 10.1128/AEM.01842-21. Epub 2021 Oct 27. Appl Environ Microbiol. 2022. PMID: 34705547 Free PMC article.
-
Fluorimetric high-throughput screening method for polyester hydrolase activity using polyethylene terephthalate nanoparticles.Methods Enzymol. 2021;648:253-270. doi: 10.1016/bs.mie.2020.11.003. Epub 2021 Jan 30. Methods Enzymol. 2021. PMID: 33579406
-
The biotechnological potential of marine bacteria in the novel lineage of Pseudomonas pertucinogena.Microb Biotechnol. 2020 Jan;13(1):19-31. doi: 10.1111/1751-7915.13288. Epub 2018 Jun 25. Microb Biotechnol. 2020. PMID: 29943398 Free PMC article. Review.
-
Synthetic polyester-hydrolyzing enzymes from thermophilic actinomycetes.Adv Appl Microbiol. 2014;89:267-305. doi: 10.1016/B978-0-12-800259-9.00007-X. Adv Appl Microbiol. 2014. PMID: 25131405 Review.
Cited by
-
XTT assay for detection of bacterial metabolic activity in water-based polyester polyurethane.PLoS One. 2024 Jun 6;19(6):e0303210. doi: 10.1371/journal.pone.0303210. eCollection 2024. PLoS One. 2024. PMID: 38843174 Free PMC article.
-
Halophilic and halotolerant fungi across diverse climates: a comparative study of Polish and Italian soil ecosystems.Front Microbiol. 2025 Jul 28;16:1637496. doi: 10.3389/fmicb.2025.1637496. eCollection 2025. Front Microbiol. 2025. PMID: 40792259 Free PMC article.
-
The Bacteroidetes Aequorivita sp. and Kaistella jeonii Produce Promiscuous Esterases With PET-Hydrolyzing Activity.Front Microbiol. 2022 Jan 5;12:803896. doi: 10.3389/fmicb.2021.803896. eCollection 2021. Front Microbiol. 2022. PMID: 35069509 Free PMC article.
-
Bioprospecting for polyesterase activity relevant for PET degradation in marine Enterobacterales isolates.AIMS Microbiol. 2023 Jun 15;9(3):518-539. doi: 10.3934/microbiol.2023027. eCollection 2023. AIMS Microbiol. 2023. PMID: 37649797 Free PMC article.
-
A High-Throughput Screening Platform for Engineering Poly(ethylene Terephthalate) Hydrolases.ACS Catal. 2024 Sep 17;14(19):14622-14638. doi: 10.1021/acscatal.4c04321. eCollection 2024 Oct 4. ACS Catal. 2024. PMID: 39386920 Free PMC article.
References
-
- Belda, E. , van Heck, R.G.A. , José Lopez‐Sanchez, M. , Cruveiller, S. , Barbe, V. , Fraser, C. , et al (2016) The revisited genome of Pseudomonas putida KT2440 enlightens its value as a robust metabolic chassis. Environ Microbiol 18: 3403–3424. - PubMed
-
- Biffinger, J.C. , Barlow, D.E. , Cockrell, A.L. , Cusick, K.D. , Hervey, W.J. , Fitzgerald, L.A. , et al (2015) The applicability of Impranil®DLN for gauging the biodegradation of polyurethanes. Polym Degrad Stab 120: 178–185.
-
- Biffinger, J.C. , Crookes‐Goodson, W.J. and Barlow, D.E. (2018) Assignment of direct vs. indirect mechanisms used by fungi for polyurethane coating degradation. SERDP Final Report for SEED WP‐2745. [WWW document] URL: https://www.serdp-estcp.org/content/download/47939/456696/file/WP-2745%2....
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

