RACK1 on and off the ribosome
- PMID: 31023766
- PMCID: PMC6573788
- DOI: 10.1261/rna.071217.119
RACK1 on and off the ribosome
Abstract
Receptor for activated C kinase 1 (RACK1) is a eukaryote-specific ribosomal protein (RP) implicated in diverse biological functions. To engineer ribosomes for specific fluorescent labeling, we selected RACK1 as a target given its location on the small ribosomal subunit and other properties. However, prior results suggested that RACK1 has roles both on and off the ribosome, and such an exchange might be related to its various cellular functions and hinder our ability to use RACK1 as a stable fluorescent tag for the ribosome. In addition, the kinetics of spontaneous exchange of RACK1 or any RP from a mature ribosome in vitro remain unclear. To address these issues, we engineered fluorescently labeled human ribosomes via RACK1, and applied bulk and single-molecule biochemical analyses to track RACK1 on and off the human ribosome. Our results demonstrate that, despite its cellular nonessentiality from yeast to humans, RACK1 readily reassociates with the ribosome, displays limited conformational dynamics, and remains stably bound to the ribosome for hours in vitro. This work sheds insight into the biochemical basis of RPs exchange on and off a mature ribosome and provides tools for single-molecule analysis of human translation.
Keywords: IRES; RACK1; human; ribosome; single-molecule fluorescence; translation.
© 2019 Johnson et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures

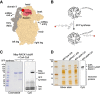


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
