Genus-wide Leptospira core genome multilocus sequence typing for strain taxonomy and global surveillance
- PMID: 31026256
- PMCID: PMC6513109
- DOI: 10.1371/journal.pntd.0007374
Genus-wide Leptospira core genome multilocus sequence typing for strain taxonomy and global surveillance
Erratum in
-
Correction: Genus-wide Leptospira core genome multilocus sequence typing for strain taxonomy and global surveillance.PLoS Negl Trop Dis. 2020 Aug 21;14(8):e0008673. doi: 10.1371/journal.pntd.0008673. eCollection 2020 Aug. PLoS Negl Trop Dis. 2020. PMID: 32822349 Free PMC article.
Abstract
Leptospira is a highly heterogeneous bacterial genus that can be divided into three evolutionary lineages and >300 serovars. The causative agents of leptospirosis are responsible of an emerging zoonotic disease worldwide. To advance our understanding of the biodiversity of Leptospira strains at the global level, we evaluated the performance of whole-genome sequencing (WGS) as a genus-wide strain classification and genotyping tool. Herein we propose a set of 545 highly conserved loci as a core genome MLST (cgMLST) genotyping scheme applicable to the entire Leptospira genus, including non-pathogenic species. Evaluation of cgMLST genotyping was undertaken with 509 genomes, including 327 newly sequenced genomes, from diverse species, sources and geographical locations. Phylogenetic analysis showed that cgMLST defines species, clades, subclades, clonal groups and cgMLST sequence types (cgST), with high precision and robustness to missing data. Novel Leptospira species, including a novel subclade named S2 (saprophytes 2), were identified. We defined clonal groups (CG) optimally using a single-linkage clustering threshold of 40 allelic mismatches. While some CGs such as L. interrogans CG6 (serogroup Icterohaemorrhagiae) are globally distributed, others are geographically restricted. cgMLST was congruent with classical MLST schemes, but had greatly improved resolution and broader applicability. Single nucleotide polymorphisms within single cgST groups was limited to <30 SNPs, underlining a potential role for cgMLST in epidemiological surveillance. Finally, cgMLST allowed identification of serogroups and closely related serovars. In conclusion, the proposed cgMLST strategy allows high-resolution genotyping of Leptospira isolates across the phylogenetic breadth of the genus. The unified genomic taxonomy of Leptospira strains, available publicly at http://bigsdb.pasteur.fr/leptospira, will facilitate global harmonization of Leptospira genotyping, strain emergence follow-up and novel collaborative studies of the epidemiology and evolution of this emerging pathogen.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

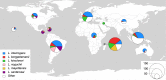

References
-
- Ellis WA. Animal Leptospirosis In: Adler B, editor. Leptospira and Leptospirosis: Springer; 2014. p. 99–137.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

