Translational Control under Stress: Reshaping the Translatome
- PMID: 31026340
- PMCID: PMC6541386
- DOI: 10.1002/bies.201900009
Translational Control under Stress: Reshaping the Translatome
Abstract
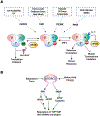
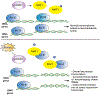
Adequate reprogramming of cellular metabolism in response to stresses or suboptimal growth conditions involves a myriad of coordinated changes that serve to promote cell survival. As protein synthesis is an energetically expensive process, its regulation under stress is of critical importance. Reprogramming of messenger RNA (mRNA) translation involves well-understood stress-activated kinases that target components of translation initiation machinery, resulting in the robust inhibition of general translation and promotion of the translation of stress-responsive proteins. Translational arrest of mRNAs also results in the accumulation of transcripts in cytoplasmic foci called stress granules. Recent studies focus on the key roles of transfer RNA (tRNA) in stress-induced translational reprogramming. These include stress-specific regulation of tRNA pools, codon-biased translation influenced by tRNA modifications, tRNA miscoding, and tRNA cleavage. In combination, signal transduction pathways and tRNA metabolism changes regulate translation during stress, resulting in adaptation and cell survival. This review examines molecular mechanisms that regulate protein synthesis in response to stress.
Keywords: stress; tRNA; translation; translation initiation; translational reprogramming.
© 2019 WILEY Periodicals, Inc.
Conflict of interest statement
Conflict of Interest
The authors declare no conflict of interest
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

