Correlation of Gut Microbiome Between ASD Children and Mothers and Potential Biomarkers for Risk Assessment
- PMID: 31026579
- PMCID: PMC6520911
- DOI: 10.1016/j.gpb.2019.01.002
Correlation of Gut Microbiome Between ASD Children and Mothers and Potential Biomarkers for Risk Assessment
Abstract
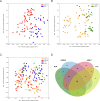
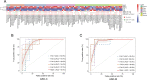
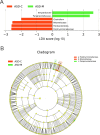
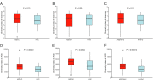
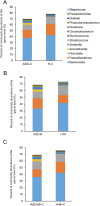
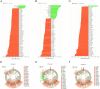
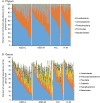
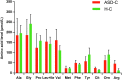
Variation of maternal gut microbiota may increase the risk of autism spectrum disorders (ASDs) in offspring. Animal studies have indicated that maternal gut microbiota is related to neurodevelopmental abnormalities in mouse offspring, while it is unclear whether there is a correlation between gut microbiota of ASD children and their mothers. We examined the relationships between gut microbiome profiles of ASD children and those of their mothers, and evaluated the clinical discriminatory power of discovered bacterial biomarkers. Gut microbiome was profiled and evaluated by 16S ribosomal RNA gene sequencing in stool samples of 59 mother-child pairs of ASD children and 30 matched mother-child pairs of healthy children. Significant differences were observed in the gut microbiome composition between ASD and healthy children in our Chinese cohort. Several unique bacterial biomarkers, such as Alcaligenaceae and Acinetobacter, were identified. Mothers of ASD children had more Proteobacteria, Alphaproteobacteria, Moraxellaceae, and Acinetobacter than mothers of healthy children. There was a clear correlation between gut microbiome profiles of children and their mothers; however, children with ASD still had unique bacterial biomarkers, such as Alcaligenaceae, Enterobacteriaceae, and Clostridium. Candidate biomarkers discovered in this study had remarkable discriminatory power. The identified patterns of mother-child gut microbiome profiles may be important for assessing risks during the early stage and planning of personalized treatment and prevention of ASD via microbiota modulation.
Keywords: Autism spectrum disorders; Biomarker; Gut microbiome; Microbiota-gut-immune-brain axis; Mother–child pair.
Copyright © 2019 The Authors. Production and hosting by Elsevier B.V. All rights reserved.
Figures


Similar articles
-
Gut Bacteria Shared by Children and Their Mothers Associate with Developmental Level and Social Deficits in Autism Spectrum Disorder.mSphere. 2020 Dec 2;5(6):e01044-20. doi: 10.1128/mSphere.01044-20. mSphere. 2020. PMID: 33268567 Free PMC article.
-
Gut Microbial Dysbiosis in Indian Children with Autism Spectrum Disorders.Microb Ecol. 2018 Nov;76(4):1102-1114. doi: 10.1007/s00248-018-1176-2. Epub 2018 Mar 21. Microb Ecol. 2018. PMID: 29564487
-
Underdevelopment of the gut microbiota and bacteria species as non-invasive markers of prediction in children with autism spectrum disorder.Gut. 2022 May;71(5):910-918. doi: 10.1136/gutjnl-2020-324015. Epub 2021 Jul 26. Gut. 2022. PMID: 34312160
-
Composition of Gut Microbiota in Children with Autism Spectrum Disorder: A Systematic Review and Meta-Analysis.Nutrients. 2020 Mar 17;12(3):792. doi: 10.3390/nu12030792. Nutrients. 2020. PMID: 32192218 Free PMC article.
-
Microbiome and nutrition in autism spectrum disorder: current knowledge and research needs.Nutr Rev. 2016 Dec;74(12):723-736. doi: 10.1093/nutrit/nuw048. Nutr Rev. 2016. PMID: 27864534 Review.
Cited by
-
Gastrointestinal symptoms have a minor impact on autism spectrum disorder and associations with gut microbiota and short-chain fatty acids.Front Microbiol. 2022 Oct 7;13:1000419. doi: 10.3389/fmicb.2022.1000419. eCollection 2022. Front Microbiol. 2022. PMID: 36274684 Free PMC article.
-
The Fast Track for Microbiome Research.Genomics Proteomics Bioinformatics. 2019 Feb;17(1):1-3. doi: 10.1016/j.gpb.2019.04.001. Epub 2019 Apr 26. Genomics Proteomics Bioinformatics. 2019. PMID: 31034986 Free PMC article. No abstract available.
-
Unraveling the immunogenetic landscape of autism spectrum disorder: a comprehensive bioinformatics approach.Front Immunol. 2024 Apr 24;15:1347139. doi: 10.3389/fimmu.2024.1347139. eCollection 2024. Front Immunol. 2024. PMID: 38726016 Free PMC article.
-
Children and adolescents with attention deficit hyperactivity disorder and autism spectrum disorder share distinct microbiota compositions.Gut Microbes. 2023 Jan-Dec;15(1):2211923. doi: 10.1080/19490976.2023.2211923. Gut Microbes. 2023. PMID: 37199526 Free PMC article.
-
Pomegranate peel extract ameliorates the severity of experimental autoimmune encephalomyelitis via modulation of gut microbiota.Gut Microbes. 2020 Nov 9;12(1):1857515. doi: 10.1080/19490976.2020.1857515. Gut Microbes. 2020. PMID: 33382357 Free PMC article.
References
-
- D'Gama A.M., Walsh C.A. Somatic mosaicism and neurodevelopmental disease. Nat Neurosci. 2018;21:1504–1514. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

