Genetically Encoded Fluorescent Indicator GRAPHIC Delineates Intercellular Connections
- PMID: 31026667
- PMCID: PMC6482341
- DOI: 10.1016/j.isci.2019.04.013
Genetically Encoded Fluorescent Indicator GRAPHIC Delineates Intercellular Connections
Abstract
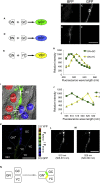
Intercellular contacts are essential for precise organ morphogenesis, function, and maintenance; however, spatiotemporal information of cell-cell contacts or adhesions remains elusive in many systems. We developed a genetically encoded fluorescent indicator for intercellular contacts with optimized intercellular GFP reconstitution using glycosylphosphatidylinositol (GPI) anchor, GRAPHIC (GPI anchored reconstitution-activated proteins highlight intercellular connections), which can be used for an expanded number of cell types. We observed a robust GFP signal specifically at the interface between cultured cells, without disrupting natural cell contact. Application of GRAPHIC to the fish retina specifically delineated cone-bipolar connection sites. Moreover, we showed that GRAPHIC can be used in the mouse central nervous system to delineate synaptic sites in the thalamocortical circuit. Finally, we generated GRAPHIC color variants, enabling detection of multiple convergent contacts simultaneously in cell culture system. We demonstrated that GRAPHIC has high sensitivity and versatility, which will facilitate the analysis of the complex multicellular connections without previous limitations.
Keywords: Biological Sciences; Cell Biology; Molecular Biology.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures





References
-
- Baes M., Denef C. Evidence that stimulation of growth hormone release by epinephrine and vasoactive intestinal peptide is based on cell-to-cell communication in the pituitary. Endocrinology. 1987;120:280–290. - PubMed
-
- Biederer T., Sudhof T.C. CASK and protein 4.1 support F-actin nucleation on neurexins. J. Biol. Chem. 2001;276:47869–47876. - PubMed
-
- Cabantous S., Terwilliger T.C., Waldo G.S. Protein tagging and detection with engineered self-assembling fragments of green fluorescent protein. Nat. Biotechnol. 2005;23:102–107. - PubMed
-
- Choi J.H., Sim S.E., Kim J.I., Choi D.I., Oh J., Ye S., Lee J., Kim T., Ko H.G., Lim C.S. Interregional synaptic maps among engram cells underlie memory formation. Science. 2018;360:430–435. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

