Modulation of the IL-6-Signaling Pathway in Liver Cells by miRNAs Targeting gp130, JAK1, and/or STAT3
- PMID: 31026677
- PMCID: PMC6479786
- DOI: 10.1016/j.omtn.2019.03.007
Modulation of the IL-6-Signaling Pathway in Liver Cells by miRNAs Targeting gp130, JAK1, and/or STAT3
Abstract
Interleukin-6 (IL-6)-type cytokines share the common receptor glycoprotein 130 (gp130), which activates a signaling cascade involving Janus kinases (JAKs) and signal transducer and activator of transcription (STAT) transcription factors. IL-6 and/or its signaling pathway is often deregulated in diseases, such as chronic liver diseases and cancer. Thus, the identification of compounds inhibiting this pathway is of interest for future targeted therapies. We established novel cellular screening systems based on a STAT-responsive reporter gene (Cypridina luciferase). Of a library containing 538 microRNA (miRNA) mimics, several miRNAs affected hyper-IL-6-induced luciferase activities. When focusing on candidate miRNAs specifically targeting 3' UTRs of signaling molecules of this pathway, we identified, e.g., miR-3677-5p as a novel miRNA affecting protein expression of both STAT3 and JAK1, whereas miR-16-1-3p, miR-4473, and miR-520f-3p reduced gp130 surface expression. Interestingly, combination treatment with 2 or 3 miRNAs targeting gp130 or different signaling molecules of the pathway did not increase the inhibitory effects on phospho-STAT3 levels and STAT3 target gene expression compared to treatment with single mimics. Taken together, we identified a set of miRNAs of potential therapeutic value for cancer and inflammatory diseases, which directly target the expression of molecules within the IL-6-signaling pathway and can dampen inflammatory signal transduction.
Keywords: JAK1; STAT3; cancer; cytokine; gp130; inflammation; interleukin-6; liver; microRNAs; signal transduction.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures



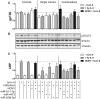



Similar articles
-
Impact of miR-223-3p and miR-2909 on inflammatory factors IL-6, IL-1ß, and TNF-α, and the TLR4/TLR2/NF-κB/STAT3 signaling pathway induced by lipopolysaccharide in human adipose stem cells.PLoS One. 2019 Feb 26;14(2):e0212063. doi: 10.1371/journal.pone.0212063. eCollection 2019. PLoS One. 2019. PMID: 30807577 Free PMC article.
-
Identification of Spinal Cord MicroRNA and Gene Signatures in a Model of Chronic Stress-Induced Visceral Hyperalgesia in Rat.PLoS One. 2015 Jul 29;10(7):e0130938. doi: 10.1371/journal.pone.0130938. eCollection 2015. PLoS One. 2015. PMID: 26222740 Free PMC article.
-
Activation of the protein tyrosine phosphatase SHP2 via the interleukin-6 signal transducing receptor protein gp130 requires tyrosine kinase Jak1 and limits acute-phase protein expression.Biochem J. 1998 Nov 1;335 ( Pt 3)(Pt 3):557-65. doi: 10.1042/bj3350557. Biochem J. 1998. PMID: 9794795 Free PMC article.
-
Interplay between microRNAs and the STAT3 signaling pathway in human cancers.Physiol Genomics. 2013 Dec 15;45(24):1206-14. doi: 10.1152/physiolgenomics.00122.2013. Epub 2013 Nov 5. Physiol Genomics. 2013. PMID: 24192393 Review.
-
Mutations leading to constitutive active gp130/JAK1/STAT3 pathway.Cytokine Growth Factor Rev. 2015 Oct;26(5):499-506. doi: 10.1016/j.cytogfr.2015.07.010. Epub 2015 Jul 3. Cytokine Growth Factor Rev. 2015. PMID: 26188635 Review.
Cited by
-
MicroRNA sequencing in patients with coronary artery disease - considerations for use as biomarker for thrombotic risk.Clin Transl Sci. 2022 Aug;15(8):1946-1958. doi: 10.1111/cts.13307. Epub 2022 May 29. Clin Transl Sci. 2022. PMID: 35643946 Free PMC article.
-
Cholangiocarcinoma.Nat Rev Dis Primers. 2021 Sep 9;7(1):65. doi: 10.1038/s41572-021-00300-2. Nat Rev Dis Primers. 2021. PMID: 34504109 Free PMC article. Review.
-
Homotherapy for Heteropathy: A Molecular Mechanism of Poria Sini Decoction for Treatment of Liver Cancer and Chronic Heart Failure.Evid Based Complement Alternat Med. 2024 Apr 29;2024:9958258. doi: 10.1155/2024/9958258. eCollection 2024. Evid Based Complement Alternat Med. 2024. PMID: 38711438 Free PMC article.
-
MiRNAs in Alcohol-Related Liver Diseases and Hepatocellular Carcinoma: A Step toward New Therapeutic Approaches?Cancers (Basel). 2023 Nov 23;15(23):5557. doi: 10.3390/cancers15235557. Cancers (Basel). 2023. PMID: 38067261 Free PMC article. Review.
-
Direct and indirect effects of IFN-α2b in malignancy treatment: not only an archer but also an arrow.Biomark Res. 2022 Sep 14;10(1):69. doi: 10.1186/s40364-022-00415-y. Biomark Res. 2022. PMID: 36104718 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

