Adaptation and Conservation throughout the Drosophila melanogaster Life-Cycle
- PMID: 31028390
- PMCID: PMC6535812
- DOI: 10.1093/gbe/evz086
Adaptation and Conservation throughout the Drosophila melanogaster Life-Cycle
Abstract
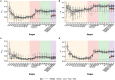
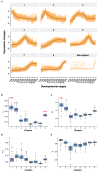
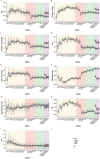
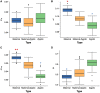
Previous studies of the evolution of genes expressed at different life-cycle stages of Drosophila melanogaster have not been able to disentangle adaptive from nonadaptive substitutions when using nonsynonymous sites. Here, we overcome this limitation by combining whole-genome polymorphism data from D. melanogaster and divergence data between D. melanogaster and Drosophila yakuba. For the set of genes expressed at different life-cycle stages of D. melanogaster, as reported in modENCODE, we estimate the ratio of substitutions relative to polymorphism between nonsynonymous and synonymous sites (α) and then α is discomposed into the ratio of adaptive (ωa) and nonadaptive (ωna) substitutions to synonymous substitutions. We find that the genes expressed in mid- and late-embryonic development are the most conserved, whereas those expressed in early development and postembryonic stages are the least conserved. Importantly, we found that low conservation in early development is due to high rates of nonadaptive substitutions (high ωna), whereas in postembryonic stages it is due, instead, to high rates of adaptive substitutions (high ωa). By using estimates of different genomic features (codon bias, average intron length, exon number, recombination rate, among others), we also find that genes expressed in mid- and late-embryonic development show the most complex architecture: they are larger, have more exons, more transcripts, and longer introns. In addition, these genes are broadly expressed among all stages. We suggest that all these genomic features are related to the conservation of mid- and late-embryonic development. Globally, our study supports the hourglass pattern of conservation and adaptation over the life-cycle.
Keywords: DFE-alpha; adaptation; conservation; evo-devo; hourglass hypothesis; natural selection.
© The Author(s) 2019. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures
References
-
- Alberch P. 1980. Ontogenesis and morphological diversification. Am Zool. 20(4):653–667.
-
- Anderson DT. 1973. Embryology and phylogeny in annelids and arthropods. Oxford, New York: Pergamon Press.
-
- Arthur W. 1977. The origin of animal body plans: a study in evolutionary developmental biology. Cambridge: Cambridge University Press.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

