Diversity of Contact-Dependent Growth Inhibition Systems of Pseudomonas aeruginosa
- PMID: 31036723
- PMCID: PMC6597396
- DOI: 10.1128/JB.00776-18
Diversity of Contact-Dependent Growth Inhibition Systems of Pseudomonas aeruginosa
Abstract
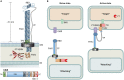
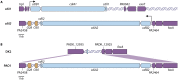
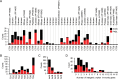
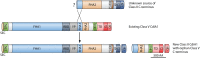
Contact-dependent growth inhibition (CDI) systems are used in bacterial competition to hinder the growth of neighboring microbes. These systems utilize a two-partner secretion mechanism to display the CdiA exoprotein at the bacterial cell surface. CdiA forms a long filamentous stalk that facilitates binding to a target cell and delivery of a C-terminal toxin (CT) domain. This CT domain is processed and delivered into the cytoplasm of a target cell upon contact. CDI systems also encode a cognate immunity protein (CdiI) that protects siblings and resistant targeted cells from intoxication by high-affinity binding to the CT. CdiA CT domains vary among strains within a species, and many alleles encode enzymatic functions that target nucleic acids. This variation is thought to help drive diversity and adaptation within a species. CdiA diversity is well studied in Escherichia coli and several other bacteria, but little is known about the extent of this diversity in Pseudomonas aeruginosa. The purpose of this review is to highlight the variability that exists in CDI systems of P. aeruginosa. We show that this diversity is apparent even among strains isolated from a single geographical region, suggesting that CDI systems play an important role in the ecology of P. aeruginosa.
Copyright © 2019 American Society for Microbiology.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

