Dynamic Network Activation of Hypothalamic MCH Neurons in REM Sleep and Exploratory Behavior
- PMID: 31036764
- PMCID: PMC6670248
- DOI: 10.1523/JNEUROSCI.0305-19.2019
Dynamic Network Activation of Hypothalamic MCH Neurons in REM Sleep and Exploratory Behavior
Abstract
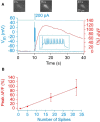
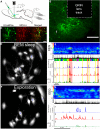
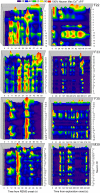
Most brain neurons are active in waking, but hypothalamic neurons that synthesize the neuropeptide melanin-concentrating hormone (MCH) are claimed to be active only during sleep, particularly rapid eye movement (REM) sleep. Here we use deep-brain imaging to identify changes in fluorescence of the genetically encoded calcium (Ca2+) indicator GCaMP6 in individual hypothalamic neurons that contain MCH. An in vitro electrophysiology study determined a strong relationship between depolarization and Ca2+ fluorescence in MCH neurons. In 10 freely behaving MCH-cre mice (male and female), the highest fluorescence occurred in all recorded neurons (n = 106) in REM sleep relative to quiet waking or non-REM sleep. Unexpectedly, 70% of the MCH neurons had strong fluorescence activity when the mice explored novel objects. Spatial and temporal mapping of the change in fluorescence between pairs of MCH neurons revealed dynamic activation of MCH neurons during REM sleep and activation of a subset of the same neurons during exploratory behavior. Functional network activity maps will facilitate comparisons of not only single-neuron activity, but also network responses in different conditions and disease.SIGNIFICANCE STATEMENT Functional activity maps identify brain circuits responding to specific behaviors, including rapid eye movement sleep (REM sleep), a sleep phase when the brain is as active as in waking. To provide the first activity map of individual neurons during REM sleep, we use deep-brain calcium imaging in unrestrained mice to map the activity of hypothalamic melanin-concentrating hormone (MCH) neurons. MCH neurons were found to be synchronously active during REM sleep, and also during the exploration of novel objects. Spatial mapping revealed dynamic network activation during REM sleep and activation of a subset of the neurons during exploratory behavior. Functional activity maps at the cellular level in specific behaviors, including sleep, are needed to establish a brain connectome.
Keywords: REM sleep; calcium imaging; hypothalamus; melanin concentrating hormone; sleep.
Copyright © 2019 the authors.
Figures


References
-
- Blanco-Centurion C, Liu M, Konadhode RP, Zhang X, Pelluru D, van den Pol AN, Shiromani PJ (2016) Optogenetic activation of melanin-concentrating hormone neurons increases non-rapid eye movement and rapid eye movement sleep during the night in rats. Eur J Neurosci 44:2846–2857. 10.1111/ejn.13410 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
