Molecular and genetic organization of bands and interbands in the dot chromosome of Drosophila melanogaster
- PMID: 31041520
- PMCID: PMC6536484
- DOI: 10.1007/s00412-019-00703-x
Molecular and genetic organization of bands and interbands in the dot chromosome of Drosophila melanogaster
Abstract
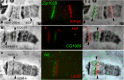
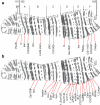
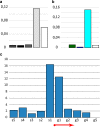
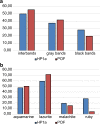
The fourth chromosome smallest in the genome of Drosophila melanogaster differs from other chromosomes in many ways. It has high repeat density in conditions of a large number of active genes. Gray bands represent a significant part of this polytene chromosome. Specific proteins including HP1a, POF, and dSETDB1 establish the epigenetic state of this unique chromatin domain. In order to compare maps of localization of genes, bands, and chromatin types of the fourth chromosome, we performed FISH analysis of 38 probes chosen according to the model of four chromatin types. It allowed clarifying the dot chromosome cytological map consisting of 16 loose gray bands, 11 dense black bands, and 26 interbands. We described the relation between chromatin states and bands. Open aquamarine chromatin mostly corresponds to interbands and it contains 5'UTRs of housekeeping genes. Their coding parts are embedded in gray bands substantially composed of lazurite chromatin of intermediate compaction. Polygenic black bands contain most of dense ruby chromatin, and also some malachite and lazurite. Having an accurate map of the fourth chromosome bands and its correspondence to physical map, we found that DNase I hypersensitivity sites, ORC2 protein, and P-elements are mainly located in open aquamarine chromatin, while element 1360, characteristic of the fourth chromosome, occupies band chromatin types. POF and HP1a proteins providing special organization of this chromosome are mostly located in aquamarine and lazurite chromatin. In general, band organization of the fourth chromosome shares the features of the whole Drosophila genome.
Keywords: Bands; Chromatin types; Dot chromosome; Drosophila; Interbands; Polytene chromosomes.
Conflict of interest statement
The authors declare that they have no conflict of interest.
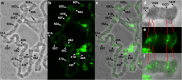
Figures









Similar articles
-
Genes Containing Long Introns Occupy Series of Bands and Interbands In Drosophila melanogaster polytene Chromosomes.Genes (Basel). 2020 Apr 11;11(4):417. doi: 10.3390/genes11040417. Genes (Basel). 2020. PMID: 32290448 Free PMC article.
-
Faint gray bands in Drosophila melanogaster polytene chromosomes are formed by coding sequences of housekeeping genes.Chromosoma. 2020 Mar;129(1):25-44. doi: 10.1007/s00412-019-00728-2. Epub 2019 Dec 9. Chromosoma. 2020. PMID: 31820086
-
CHARACTERISTIC OF THE CHROMATIN TYPE CORRESPONDING TO THIN «GREY» BANDS IN POLYTHENE CHROMOSOMES OF DROSOPHILA MELANOGASTER.Tsitologiia. 2016;58(4):248-52. Tsitologiia. 2016. PMID: 30191688 English, Russian.
-
[Chromomeric organization of interphase chromosomes in Drosophila melanogaster].Tsitologiia. 2013;55(3):144-7. Tsitologiia. 2013. PMID: 23795454 Review. Russian.
-
Polytene Chromosomes - A Portrait of Functional Organization of the Drosophila Genome.Curr Genomics. 2018 Apr;19(3):179-191. doi: 10.2174/1389202918666171016123830. Curr Genomics. 2018. PMID: 29606905 Free PMC article. Review.
Cited by
-
Genes Containing Long Introns Occupy Series of Bands and Interbands In Drosophila melanogaster polytene Chromosomes.Genes (Basel). 2020 Apr 11;11(4):417. doi: 10.3390/genes11040417. Genes (Basel). 2020. PMID: 32290448 Free PMC article.
-
Interchromosomal Contacts of rDNA Clusters in Three Human Cell Lines Are Associated with Silencing of Genes Controlling Morphogenesis.Dokl Biochem Biophys. 2021 May;496(1):22-26. doi: 10.1134/S1607672921010038. Epub 2021 Mar 10. Dokl Biochem Biophys. 2021. PMID: 33689069
-
The Organization of Pericentromeric Heterochromatin in Polytene Chromosome 3 of the Drosophilamelanogaster Line with the Rif11; SuURES Su(var)3-906 Mutations Suppressing Underreplication.Cells. 2021 Oct 20;10(11):2809. doi: 10.3390/cells10112809. Cells. 2021. PMID: 34831030 Free PMC article.
-
Effects of Mutations in the Drosophila melanogaster Rif1 Gene on the Replication and Underreplication of Pericentromeric Heterochromatin in Salivary Gland Polytene Chromosomes.Cells. 2020 Jun 19;9(6):1501. doi: 10.3390/cells9061501. Cells. 2020. PMID: 32575592 Free PMC article.
References
-
- Andreyeva EN, Kolesnikova TD, Demakova OV, Mendez-Lago M, Pokholkova GV, Belyaeva ES, Rossi F, Dimitri P, Villasante A, Zhimulev IF. High-resolution analysis of Drosophila heterochromatin organization using SuUR Su(var)3-9 double mutants. Proc Natl Acad Sci U S A. 2007;104:12819–12824. doi: 10.1073/pnas.0704690104. - DOI - PMC - PubMed
-
- Ashburner M, Golic KG, Hawley RS. Drosophila: a laboratory handbook. 2. Cold Spring Harbor: Cold Spring Harbor Laboratory Press; 2005.
-
- Boldyreva LV, Goncharov FP, Demakova OV, Zykova TY, Levitsky VG, Kolesnikov NN, Pindyurin AV, Semeshin VF, Zhimulev IF. Protein and genetic composition of four chromatin types in Drosophila melanogaster cell lines. Curr Genomics. 2017;18:214–226. doi: 10.2174/1389202917666160512164913. - DOI - PMC - PubMed
-
- Bridges CB. Salivary chromosome maps with a key to the banding of the chromosomes of Drosophila melanogaster. J Hered. 1935;26:60–64. doi: 10.1093/oxfordjournals.jhered.a104022. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

