Mechanism of the Flavoprotein d-6-Hydroxynicotine Oxidase: Substrate Specificity, pH and Solvent Isotope Effects, and Roles of Key Active-Site Residues
- PMID: 31046245
- PMCID: PMC6786761
- DOI: 10.1021/acs.biochem.9b00297
Mechanism of the Flavoprotein d-6-Hydroxynicotine Oxidase: Substrate Specificity, pH and Solvent Isotope Effects, and Roles of Key Active-Site Residues
Abstract
The flavoprotein d-6-hydroxynicotine oxidase catalyzes an early step in the oxidation of ( R)-nicotine, the oxidation of a carbon-nitrogen bond in the pyrrolidine ring of ( R)-6-hydroxynicotine. The enzyme is a member of the vanillyl alcohol oxidase/ p-cresol methylhydroxylase family of flavoproteins. The effects of substrate modifications on the steady-state and rapid-reaction kinetic parameters are not consistent with the quinone-methide mechanism of p-cresol methylhydroxylase. There is no solvent isotope effect on the kcat/ Kamine value with either ( R)-6-hydroxynicotine or the slower substrate ( R)-6-hydroxynornicotine. The effect of pH on the rapid-reaction kinetic parameters establishes that only the neutral form of the substrate and the correctly protonated form of the enzyme bind. The active-site residues Lys348, Glu350, and Glu352 are all properly positioned for substrate binding. The K348M substitution has only a small effect on the kinetic parameters; the E350A and E350Q substitutions decrease the kcat/ Kamine value by ∼20- and ∼220-fold, respectively, and the E352Q substitution decreases this parameter ∼3800-fold. The kcat/ Kamine-pH profile is bell-shaped. The p Ka values in that profile are altered by replacement of ( R)-6-hydroxynicotine with ( R)-6-hydroxynornicotine as the substrate and by the substitutions for Glu350 and Glu352, although the profiles remain bell-shaped. The results are consistent with a network of hydrogen-bonded residues in the active site being involved in binding the neutral form of the amine substrate, followed by the transfer of a hydride from the amine to the flavin.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
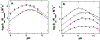
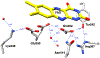
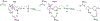
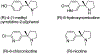


References
-
- Brandsch R (2006) Microbiology and biochemistry of nicotine degradation. Appl. Microbiol. Biotechnol 69, 493–498. - PubMed
-
- Wang SN, Liu Z, Tang HZ, Meng J, and Xu P (2007) Characterization of environmentally friendly nicotine degradation by Pseudomonas putida biotype A strain S16. Microbiology 153, 1556–1565. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

