Post-Translational Modifications of the Mini-Chromosome Maintenance Proteins in DNA Replication
- PMID: 31052337
- PMCID: PMC6563057
- DOI: 10.3390/genes10050331
Post-Translational Modifications of the Mini-Chromosome Maintenance Proteins in DNA Replication
Abstract
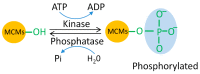
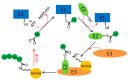
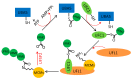
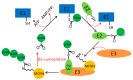
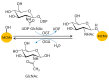
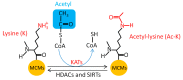
The eukaryotic mini-chromosome maintenance (MCM) complex, composed of MCM proteins 2-7, is the core component of the replisome that acts as the DNA replicative helicase to unwind duplex DNA and initiate DNA replication. MCM10 tightly binds the cell division control protein 45 homolog (CDC45)/MCM2-7/ DNA replication complex Go-Ichi-Ni-San (GINS) (CMG) complex that stimulates CMG helicase activity. The MCM8-MCM9 complex may have a non-essential role in activating the pre-replicative complex in the gap 1 (G1) phase by recruiting cell division cycle 6 (CDC6) to the origin recognition complex (ORC). Each MCM subunit has a distinct function achieved by differential post-translational modifications (PTMs) in both DNA replication process and response to replication stress. Such PTMs include phosphorylation, ubiquitination, small ubiquitin-like modifier (SUMO)ylation, O-N-acetyl-D-glucosamine (GlcNAc)ylation, and acetylation. These PTMs have an important role in controlling replication progress and genome stability. Because MCM proteins are associated with various human diseases, they are regarded as potential targets for therapeutic development. In this review, we summarize the different PTMs of the MCM proteins, their involvement in DNA replication and disease development, and the potential therapeutic implications.
Keywords: DNA replication; cancer; cell cycle; disease; genome stability; mini-chromosome maintenance (MCM); post-translational modifications (PTMs).
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
The Cdc45·Mcm2-7·GINS protein complex in trypanosomes regulates DNA replication and interacts with two Orc1-like proteins in the origin recognition complex.J Biol Chem. 2011 Sep 16;286(37):32424-35. doi: 10.1074/jbc.M111.240143. Epub 2011 Jul 28. J Biol Chem. 2011. PMID: 21799014 Free PMC article.
-
Site-specific MCM sumoylation prevents genome rearrangements by controlling origin-bound MCM.PLoS Genet. 2022 Jun 13;18(6):e1010275. doi: 10.1371/journal.pgen.1010275. eCollection 2022 Jun. PLoS Genet. 2022. PMID: 35696436 Free PMC article.
-
Termination of DNA replication forks: "Breaking up is hard to do".Nucleus. 2015;6(3):187-96. doi: 10.1080/19491034.2015.1035843. Epub 2015 Apr 2. Nucleus. 2015. PMID: 25835602 Free PMC article.
-
Regulation of Cdc45 in the cell cycle and after DNA damage.Biochem Soc Trans. 2009 Aug;37(Pt 4):926-30. doi: 10.1042/BST0370926. Biochem Soc Trans. 2009. PMID: 19614620 Review.
-
Behavior of replication origins in Eukaryota - spatio-temporal dynamics of licensing and firing.Cell Cycle. 2015;14(14):2251-64. doi: 10.1080/15384101.2015.1056421. Epub 2015 Jun 1. Cell Cycle. 2015. PMID: 26030591 Free PMC article. Review.
Cited by
-
CDK9-55 guides the anaphase-promoting complex/cyclosome (APC/C) in choosing the DNA repair pathway choice.Oncogene. 2024 Apr;43(17):1263-1273. doi: 10.1038/s41388-024-02982-w. Epub 2024 Mar 4. Oncogene. 2024. PMID: 38433256
-
DONSON and FANCM associate with different replisomes distinguished by replication timing and chromatin domain.Nat Commun. 2020 Aug 7;11(1):3951. doi: 10.1038/s41467-020-17449-1. Nat Commun. 2020. PMID: 32769987 Free PMC article.
-
Regulation of EBNA1 protein stability and DNA replication activity by PLOD1 lysine hydroxylase.PLoS Pathog. 2023 Jun 1;19(6):e1010478. doi: 10.1371/journal.ppat.1010478. eCollection 2023 Jun. PLoS Pathog. 2023. PMID: 37262099 Free PMC article.
-
MCM proteins are up-regulated in placentas of women with reduced insulin sensitivity.Biosci Rep. 2024 Oct 30;44(10):BSR20240430. doi: 10.1042/BSR20240430. Biosci Rep. 2024. PMID: 39268985 Free PMC article.
-
Unwinding Helicase MCM Functionality for Diagnosis and Therapeutics of Replication Abnormalities Associated with Cancer: A Review.Mol Diagn Ther. 2024 May;28(3):249-264. doi: 10.1007/s40291-024-00701-5. Epub 2024 Mar 26. Mol Diagn Ther. 2024. PMID: 38530633 Review.
References
-
- D’Angiolella V., Donato V., Forrester F.M., Jeong Y.T., Pellacani C., Kudo Y., Saraf A., Florens L., Washburn M.P., Pagano M. Cyclin F-mediated degradation of ribonucleotide reductase M2 controls genome integrity and DNA repair. Cell. 2012;149:1023–1034. doi: 10.1016/j.cell.2012.03.043. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

