Structure-guided post-SELEX optimization of an ochratoxin A aptamer
- PMID: 31062016
- PMCID: PMC6582339
- DOI: 10.1093/nar/gkz336
Structure-guided post-SELEX optimization of an ochratoxin A aptamer
Abstract
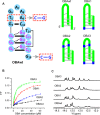
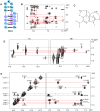
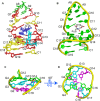
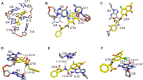
SELEX is the cornerstone for aptamer research with broad applications in biosensors and medicine. To improve the affinity of selected aptamers, we propose a structure-guided post-SELEX approach, an optimization method based on the precise secondary structure of the aptamer-ligand complex. We demonstrate this approach using the Ochratoxin A (OTA) aptamer. Guided by the structure, we designed a new aptamer whose affinity is improved by more than 50-fold. We also determined the high-resolution NMR structure of the new aptamer-OTA complex and elucidated the discriminatory recognition mechanism of one atomic difference between two analogs, OTA and OTB. The aptamer forms an unusual hairpin structure containing an intramolecular triple helix, which is not seen in the previously determined aptamer complex. The π-π stacking, the hydrophobic interaction, hydrogen bonds and halogen bonds between OTA and the aptamer contribute to the recognition of OTA, and the halogen bonds play an important role in discriminating between OTA and OTB. Our results demonstrate that the structure-guided post-SELEX approach improves aptamers affinity. An improved OTA biosensor system might be developed using this new strategy.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


References
-
- Tuerk C., Gold L.. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science. 1990; 249:505–510. - PubMed
-
- Ellington A.D., Szostak J.W.. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990; 346:818–822. - PubMed
-
- Verdian A. Apta-nanosensors for detection and quantitative determination of acetamiprid - a pesticide residue in food and environment. Talanta. 2018; 176:456–464. - PubMed
-
- Syed M.A., Jamil B.. Aptamers and aptasensors as novel approach for microbial detection and identification: an appraisal. Curr. Drug Targets. 2018; 19:1560–1572. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

