Genetic Variation for Seed Metabolite Levels in Brachypodium distachyon
- PMID: 31083584
- PMCID: PMC6540107
- DOI: 10.3390/ijms20092348
Genetic Variation for Seed Metabolite Levels in Brachypodium distachyon
Abstract
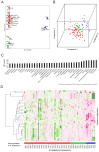
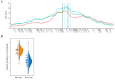
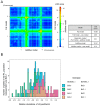
Metabolite composition and concentrations in seed grains are important traits of cereals. To identify the variation in the seed metabolotypes of a model grass, namely Brachypodium distachyon, we applied a widely targeted metabolome analysis to forty inbred lines of B. distachyon and examined the accumulation patterns of 183 compounds in the seeds. By comparing the metabolotypes with the population structure of these lines, we found signature metabolites that represent different accumulation patterns for each of the three B. distachyon subpopulations. Moreover, we found that thirty-seven metabolites exhibited significant differences in their accumulation between the lines Bd21 and Bd3-1. Using a recombinant inbred line (RIL) population from a cross between Bd3-1 and Bd21, we identified the quantitative trait loci (QTLs) linked with this variation in the accumulation of thirteen metabolites. Our metabolite QTL analysis illustrated that different genetic factors may presumably regulate the accumulation of 4-pyridoxate and pyridoxamine in vitamin B6 metabolism. Moreover, we found two QTLs on chromosomes 1 and 4 that affect the accumulation of an anthocyanin, chrysanthemin. These QTLs genetically interacted to regulate the accumulation of this compound. This study demonstrates the potential for metabolite QTL mapping in B. distachyon and provides new insights into the genetic dissection of metabolomic traits in temperate grasses.
Keywords: Brachypodium distachyon; QTL analysis; chrysanthemin; metabolome; recombinant inbred line; vitamin B6.
Conflict of interest statement
The authors declare no conflict of interest. Mention of trade names or commercial products in this publication is solely for providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture (USDA). USDA is an equal opportunity provider and employer. The complete nondiscrimination policy can be found on the USDA website.
Figures


References
-
- Bruinsma J. World Agriculture: Towards 2015/2030. Summary Report. Food and Agriculture Organization of the United Nations; Rome Italy: 2002.
-
- McKevith B. Nutritional aspects of cereals. Nutr. Bulletin. 2004;29:111–142. doi: 10.1111/j.1467-3010.2004.00418.x. - DOI
-
- Harrigan G.G., Martino-Catt S., Glenn K.C. Metabolomics, metabolic diversity and genetic variation in crops. Metabolomics. 2007;3:259–272. doi: 10.1007/s11306-007-0076-0. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

