Copy number variant and runs of homozygosity detection by microarrays enabled more precise molecular diagnoses in 11,020 clinical exome cases
- PMID: 31101064
- PMCID: PMC6525387
- DOI: 10.1186/s13073-019-0639-5
Copy number variant and runs of homozygosity detection by microarrays enabled more precise molecular diagnoses in 11,020 clinical exome cases
Abstract
Background: Exome sequencing (ES) has been successfully applied in clinical detection of single nucleotide variants (SNVs) and small indels. However, identification of copy number variants (CNVs) using ES data remains challenging. The purpose of this study is to understand the contribution of CNVs and copy neutral runs of homozygosity (ROH) in molecular diagnosis of patients referred for ES.
Methods: In a cohort of 11,020 consecutive ES patients, an Illumina SNP array analysis interrogating mostly coding SNPs was performed as a quality control (QC) measurement and for CNV/ROH detection. Among these patients, clinical chromosomal microarray analysis (CMA) was performed at Baylor Genetics (BG) on 3229 patients, either before, concurrently, or after ES. We retrospectively analyzed the findings from CMA and the QC array.
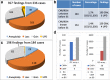
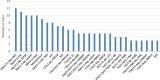
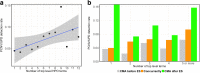
Results: The QC array can detect ~ 70% of pathogenic/likely pathogenic CNVs (PCNVs) detectable by CMA. Out of the 11,020 ES cases, the QC array identified PCNVs in 327 patients and uniparental disomy (UPD) disorder-related ROH in 10 patients. The overall PCNV/UPD detection rate was 5.9% in the 3229 ES patients who also had CMA at BG; PCNV/UPD detection rate was higher in concurrent ES and CMA than in ES with prior CMA (7.2% vs 4.6%). The PCNVs/UPD contributed to the molecular diagnoses in 17.4% (189/1089) of molecularly diagnosed ES cases with CMA and were estimated to contribute in 10.6% of all molecularly diagnosed ES cases. Dual diagnoses with both PCNVs and SNVs were detected in 38 patients. PCNVs affecting single recessive disorder genes in a compound heterozygous state with SNVs were detected in 4 patients, and homozygous deletions (mostly exonic deletions) were detected in 17 patients. A higher PCNV detection rate was observed for patients with syndromic phenotypes and/or cardiovascular abnormalities.
Conclusions: Our clinical genomics study demonstrates that detection of PCNV/UPD through the QC array or CMA increases ES diagnostic rate, provides more precise molecular diagnosis for dominant as well as recessive traits, and enables more complete genetic diagnoses in patients with dual or multiple molecular diagnoses. Concurrent ES and CMA using an array with exonic coverage for disease genes enables most effective detection of both CNVs and SNVs and therefore is recommended especially in time-sensitive clinical situations.
Keywords: Dual molecular diagnoses; Exome sequencing; Exonic CNV in AR disorders; Microarray; ROH; Structural variation; Uniparental disomy.
Conflict of interest statement
Baylor College of Medicine and Miraca Holdings Inc. have formed a joint venture with shared ownership and governance of Baylor Genetics (BG), formerly the Baylor Miraca Genetics Laboratories, which performs chromosomal microarray analysis and clinical exome sequencing. JRL serves on the Scientific Advisory Board of the BG. JRL has stock ownership in 23andMe, is a paid consultant for Regeneron Pharmaceuticals, and is a co-inventor on multiple US and European patents related to molecular diagnostics for inherited neuropathies, eye diseases, and bacterial genomic fingerprinting. Yang is a member of the Scientific Advisory Board (SAB) of Veritas Genetics China. The remaining authors declare that they have no competing interests.
Figures

Similar articles
-
Simultaneous Detection of CNVs and SNVs Improves the Diagnostic Yield of Fetuses with Ultrasound Anomalies and Normal Karyotypes.Genes (Basel). 2020 Nov 25;11(12):1397. doi: 10.3390/genes11121397. Genes (Basel). 2020. PMID: 33255631 Free PMC article.
-
Clinical utility of regions of homozygosity (ROH) identified in exome sequencing: when to pursue confirmatory uniparental disomy testing for imprinting disorders?Clin Chem Lab Med. 2024 Jul 19;63(1):87-96. doi: 10.1515/cclm-2024-0239. Print 2025 Jan 29. Clin Chem Lab Med. 2024. PMID: 39022805
-
Efficient detection of chromosome imbalances and single nucleotide variants using targeted sequencing in the clinical setting.Eur J Med Genet. 2017 Dec;60(12):667-674. doi: 10.1016/j.ejmg.2017.08.020. Epub 2017 Sep 4. Eur J Med Genet. 2017. PMID: 28882788
-
Molecular Diagnostic Yield of Exome Sequencing and Chromosomal Microarray in Cerebral Palsy: A Systematic Review and Meta-analysis.JAMA Neurol. 2022 Dec 1;79(12):1287-1295. doi: 10.1001/jamaneurol.2022.3549. JAMA Neurol. 2022. PMID: 36279113 Free PMC article.
-
Prenatal diagnosis in the fetal hyperechogenic kidneys: assessment using chromosomal microarray analysis and exome sequencing.Hum Genet. 2023 Jun;142(6):835-847. doi: 10.1007/s00439-023-02545-1. Epub 2023 Apr 24. Hum Genet. 2023. PMID: 37095353 Review.
Cited by
-
Clinical genomics and contextualizing genome variation in the diagnostic laboratory.Expert Rev Mol Diagn. 2020 Oct;20(10):995-1002. doi: 10.1080/14737159.2020.1826312. Epub 2020 Oct 10. Expert Rev Mol Diagn. 2020. PMID: 32954863 Free PMC article. Review.
-
Concordance Between Biochemical and Molecular Diagnosis Obtained by WES in Mexican Patients with Inborn Errors of Intermediary Metabolism: Utility for Therapeutic Management.Int J Mol Sci. 2024 Oct 31;25(21):11722. doi: 10.3390/ijms252111722. Int J Mol Sci. 2024. PMID: 39519275 Free PMC article.
-
Integrated sequencing and array comparative genomic hybridization in familial Parkinson disease.Neurol Genet. 2020 Jul 28;6(5):e498. doi: 10.1212/NXG.0000000000000498. eCollection 2020 Oct. Neurol Genet. 2020. PMID: 32802956 Free PMC article.
-
The diagnostic yield of CGH and WES in neurodevelopmental disorders.Front Pediatr. 2023 Mar 1;11:1133789. doi: 10.3389/fped.2023.1133789. eCollection 2023. Front Pediatr. 2023. PMID: 36937954 Free PMC article.
-
The utility of DNA methylation signatures in directing genome sequencing workflow: Kabuki syndrome and CDK13-related disorder.Am J Med Genet A. 2022 May;188(5):1368-1375. doi: 10.1002/ajmg.a.62650. Epub 2022 Jan 18. Am J Med Genet A. 2022. PMID: 35043535 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

