Quantification of Microbial Source Tracking and Pathogenic Bacterial Markers in Water and Sediments of Tiaoxi River (Taihu Watershed)
- PMID: 31105648
- PMCID: PMC6492492
- DOI: 10.3389/fmicb.2019.00699
Quantification of Microbial Source Tracking and Pathogenic Bacterial Markers in Water and Sediments of Tiaoxi River (Taihu Watershed)
Abstract
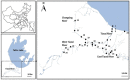
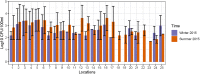
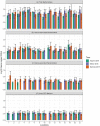
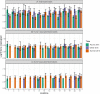
Taihu Lake is one of the largest freshwater lakes in China, serving as an important source of drinking water; >60% of source water to this lake is provided by the Tiaoxi River. This river faces serious fecal contamination issues, and therefore, a comprehensive investigation to identify the sources of fecal contamination was carried out and is presented here. The performance of existing universal (BacUni and GenBac), human (HF183-Taqman, HF183-SYBR, BacHum, and Hum2), swine (Pig-2-Bac), ruminant (BacCow), and avian (AV4143 and GFD) associated microbial source tracking (MST) markers was evaluated prior to their application in this region. The specificity and sensitivity results indicated that BacUni, HF183-TaqMan, Pig-2-Bac, and GFD assays are the most suitable in identifying human and animal fecal contamination. Therefore, these markers along with marker genes specific to selected bacterial pathogens were quantified in water and sediment samples of the Tiaoxi River, collected from 15 locations over three seasons during 2014 and 2015. Total/universal Bacteroidales markers were detected in all water and sediment samples (mean concentration 6.22 log10 gene copies/100 ml and 6.11 log10 gene copies/gram, respectively), however, the detection of host-associated MST markers varied. Human and avian markers were the most frequently detected in water samples (97 and 89%, respectively), whereas in sediment samples, only human-associated markers were detected more often (86%) than swine (64%) and avian (8.8%) markers. The results indicate that several locations in the Tiaoxi River are heavily polluted by fecal contamination and this correlated well with land use patterns. Among the five bacterial pathogens tested, Shigella spp. and Campylobacter jejuni were the most frequently detected pathogens in water (60% and 62%, respectively) and sediment samples (91% and 53%, respectively). Shiga toxin-producing Escherichia coli (STEC) and pathogenic Leptospira spp. were less frequently detected in water samples (55% and 33%, respectively) and sediment samples (51% and 13%, respectively), whereas E. coli O157:H7 was only detected in sediment samples (11%). Overall, the higher prevalence and concentrations of Campylobacter jejuni, Shigella spp., and STEC, along with the MST marker detection at a number of locations in the Tiaoxi River, indicates poor water quality and a significant human health risk associated with this watercourse. GRAPHICAL ABSTRACTTracking fecal contamination and pathogens in watersheds using molecular methods.
Keywords: Taihu watershed; Tiaoxi River; fecal pollution; microbial source tracking (MST); pathogens; qPCR quantification.
Figures

Similar articles
-
Synergistic Application of Molecular Markers and Community-Based Microbial Source Tracking Methods for Identification of Fecal Pollution in River Water During Dry and Wet Seasons.Front Microbiol. 2021 Jun 14;12:660368. doi: 10.3389/fmicb.2021.660368. eCollection 2021. Front Microbiol. 2021. PMID: 34194406 Free PMC article.
-
Microbial source tracking (MST) in Chattahoochee River National Recreation Area: Seasonal and precipitation trends in MST marker concentrations, and associations with E. coli levels, pathogenic marker presence, and land use.Water Res. 2020 Mar 15;171:115435. doi: 10.1016/j.watres.2019.115435. Epub 2019 Dec 26. Water Res. 2020. PMID: 31927096 Free PMC article.
-
Next-generation sequencing reveals fecal contamination and potentially pathogenic bacteria in a major inflow river of Taihu Lake.Environ Pollut. 2019 Nov;254(Pt B):113108. doi: 10.1016/j.envpol.2019.113108. Epub 2019 Aug 28. Environ Pollut. 2019. PMID: 31491696
-
Evaluation of host-specific Bacteroidales 16S rRNA gene markers as a complementary tool for detecting fecal pollution in a prairie watershed.Water Res. 2009 Nov;43(19):4838-49. doi: 10.1016/j.watres.2009.06.045. Epub 2009 Jun 27. Water Res. 2009. PMID: 19604534
-
Beyond borders: A systematic review and meta-analysis of human-specific faecal markers across geographical settings.Crit Rev Environ Sci Technol. 2025 Feb 6;55(7):447-464. doi: 10.1080/10643389.2025.2455031. eCollection 2025. Crit Rev Environ Sci Technol. 2025. PMID: 40329995 Free PMC article. Review.
Cited by
-
An overview of molecular markers for identification of non-human fecal pollution sources.Front Microbiol. 2023 Nov 23;14:1256174. doi: 10.3389/fmicb.2023.1256174. eCollection 2023. Front Microbiol. 2023. PMID: 38075863 Free PMC article. Review.
-
Waterborne pathogens detection technologies: advances, challenges, and future perspectives.Front Microbiol. 2023 Nov 23;14:1286923. doi: 10.3389/fmicb.2023.1286923. eCollection 2023. Front Microbiol. 2023. PMID: 38075917 Free PMC article. Review.
-
Validation of microbial source tracking markers for the attribution of fecal contamination in indoor-household environments of the Peruvian Amazon.Sci Total Environ. 2020 Nov 15;743:140531. doi: 10.1016/j.scitotenv.2020.140531. Epub 2020 Jul 2. Sci Total Environ. 2020. PMID: 32758812 Free PMC article.
-
Synergistic Application of Molecular Markers and Community-Based Microbial Source Tracking Methods for Identification of Fecal Pollution in River Water During Dry and Wet Seasons.Front Microbiol. 2021 Jun 14;12:660368. doi: 10.3389/fmicb.2021.660368. eCollection 2021. Front Microbiol. 2021. PMID: 34194406 Free PMC article.
-
Comparative Analysis of Fecal Microbiomes From Wild Waterbirds to Poultry, Cattle, Pigs, and Wastewater Treatment Plants for a Microbial Source Tracking Approach.Front Microbiol. 2021 Jul 14;12:697553. doi: 10.3389/fmicb.2021.697553. eCollection 2021. Front Microbiol. 2021. PMID: 34335529 Free PMC article.
References
-
- Ahmed W., Sritharan T., Palmer A., Sidhu J. P., Toze S. (2013). Evaluation of bovine feces-associated microbial source tracking markers and their correlations with fecal indicators and zoonotic pathogens in a Brisbane, Australia, reservoir. Appl. Environ. Microbiol. 79 2682–2691. 10.1128/AEM.03234-12 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

