Impact of Warming on Greenhouse Gas Production and Microbial Diversity in Anoxic Peat From a Sphagnum-Dominated Bog (Grand Rapids, Minnesota, United States)
- PMID: 31105668
- PMCID: PMC6498409
- DOI: 10.3389/fmicb.2019.00870
Impact of Warming on Greenhouse Gas Production and Microbial Diversity in Anoxic Peat From a Sphagnum-Dominated Bog (Grand Rapids, Minnesota, United States)
Abstract
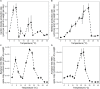
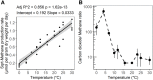
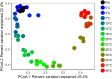
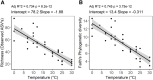
Climate warming is predicted to increase heterotrophic metabolism in northern peatland soils leading to enhanced greenhouse gas emissions. However, the specific relationships between temperature and the greenhouse gas producing microbial communities are poorly understood. Thus, in this study, the temperature dependence of carbon dioxide (CO2) and methane (CH4) production rates along with abundance and composition of microbial communities were investigated in peat from a Sphagnum-dominated peatland, S1 bog (Minnesota, United States). Whereas CH4 production rates increased with temperature up to 30°C, CO2 production did not, resulting in a lower CO2:CH4 ratio with increasing temperature. CO2 production showed both psychrophilic and mesophilic maxima at 4 and 20°C, respectively, and appears to be mediated by two anaerobic microbial communities, one that operates under psychrophilic conditions that predominate for much of the year, and another that is more active under warmer conditions during the growing season. In incubations at 10°C above the ambient range, members of the Clostridiaceae and hydrogenotrophic methanogens of the Methanobacteriaceae dominated. Moreover, a significant negative correlation between temperature and microbial diversity was observed. Results indicate that the potential consequences of warming surface peat in northern peatlands include a large stimulation in CH4 production and a significant loss of microbial diversity.
Keywords: climate change; methanogenesis; microbial community; microbial diversity; peatlands.
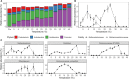
Figures
Similar articles
-
Deep peat warming increases surface methane and carbon dioxide emissions in a black spruce-dominated ombrotrophic bog.Glob Chang Biol. 2017 Dec;23(12):5398-5411. doi: 10.1111/gcb.13806. Epub 2017 Jul 28. Glob Chang Biol. 2017. PMID: 28675635
-
Porewater constituents inhibit microbially mediated greenhouse gas production (GHG) and regulate the response of soil organic matter decomposition to warming in anoxic peat from a Sphagnum-dominated bog.FEMS Microbiol Ecol. 2023 Jun 16;99(7):fiad060. doi: 10.1093/femsec/fiad060. FEMS Microbiol Ecol. 2023. PMID: 37280172
-
The role of oxygen in stimulating methane production in wetlands.Glob Chang Biol. 2021 Nov;27(22):5831-5847. doi: 10.1111/gcb.15831. Epub 2021 Aug 18. Glob Chang Biol. 2021. PMID: 34409684 Free PMC article.
-
Methane production and oxidation potentials along a fen-bog gradient from southern boreal to subarctic peatlands in Finland.Glob Chang Biol. 2021 Sep;27(18):4449-4464. doi: 10.1111/gcb.15740. Epub 2021 Jun 28. Glob Chang Biol. 2021. PMID: 34091981
-
Biogeochemical transformation of greenhouse gas emissions from terrestrial to atmospheric environment and potential feedback to climate forcing.Environ Sci Pollut Res Int. 2020 Nov;27(31):38513-38536. doi: 10.1007/s11356-020-10151-1. Epub 2020 Aug 8. Environ Sci Pollut Res Int. 2020. PMID: 32770337 Review.
Cited by
-
Autumn destabilization of deep porewater CO2 store in a northern peatland driven by turbulent diffusion.Nat Commun. 2021 Nov 25;12(1):6857. doi: 10.1038/s41467-021-27059-0. Nat Commun. 2021. PMID: 34824219 Free PMC article.
-
Soil metabolome response to whole-ecosystem warming at the Spruce and Peatland Responses under Changing Environments experiment.Proc Natl Acad Sci U S A. 2021 Jun 22;118(25):e2004192118. doi: 10.1073/pnas.2004192118. Proc Natl Acad Sci U S A. 2021. PMID: 34161254 Free PMC article.
-
Differential Impacts of Water Table and Temperature on Bacterial Communities in Pore Water From a Subalpine Peatland, Central China.Front Microbiol. 2021 May 28;12:649981. doi: 10.3389/fmicb.2021.649981. eCollection 2021. Front Microbiol. 2021. PMID: 34122363 Free PMC article.
-
Experimental warming alters the community composition, diversity, and N2 fixation activity of peat moss (Sphagnum fallax) microbiomes.Glob Chang Biol. 2019 Sep;25(9):2993-3004. doi: 10.1111/gcb.14715. Epub 2019 Jul 2. Glob Chang Biol. 2019. PMID: 31148286 Free PMC article.
-
Functional capacities of microbial communities to carry out large scale geochemical processes are maintained during ex situ anaerobic incubation.PLoS One. 2021 Feb 25;16(2):e0245857. doi: 10.1371/journal.pone.0245857. eCollection 2021. PLoS One. 2021. PMID: 33630888 Free PMC article.
References
-
- Allison S. D., Wallenstein M. D., Bradford M. A. (2010). Soil-carbon response to warming dependent on microbial physiology. Nat. Geosci. 3 336–340. 10.1038/ngeo846 - DOI
-
- Bižić-Ionescu M., Klintzch T., Ionescu D., Hindiyeh M. Y., Guenthel M., Muro Pastor A. M., et al. (2018). Widespread formation of methane by Cyanobacteria in aquatic and terrestrial environments. bioRxiv [Preprint] 10.1101/398958 - DOI
LinkOut - more resources
Full Text Sources

