The CXCR4-LASP1-eIF4F Axis Promotes Translation of Oncogenic Proteins in Triple-Negative Breast Cancer Cells
- PMID: 31106142
- PMCID: PMC6499106
- DOI: 10.3389/fonc.2019.00284
The CXCR4-LASP1-eIF4F Axis Promotes Translation of Oncogenic Proteins in Triple-Negative Breast Cancer Cells
Abstract
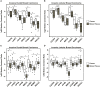
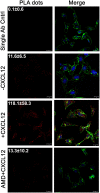
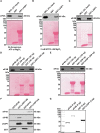
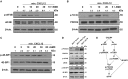
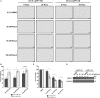
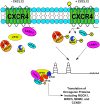
Triple-negative breast cancer (TNBC) remains clinically challenging as effective targeted therapies are lacking. In addition, patient mortality mainly results from the metastasized lesions. CXCR4 has been identified to be one of the major chemokine receptors involved in breast cancer metastasis. Previously, our lab had identified LIM and SH3 Protein 1 (LASP1) to be a key mediator in CXCR4-driven invasion. To further investigate the role of LASP1 in this process, a proteomic screen was employed and identified a novel protein-protein interaction between LASP1 and components of eukaryotic initiation 4F complex (eIF4F). We hypothesized that activation of the CXCR4-LASP1-eIF4F axis may contribute to the preferential translation of oncogenic mRNAs leading to breast cancer progression and metastasis. To test this hypothesis, we first confirmed that the gene expression of CXCR4, LASP1, and eIF4A are upregulated in invasive breast cancer. Moreover, we demonstrate that LASP1 associated with eIF4A in a CXCL12-dependent manner via a proximity ligation assay. We then confirmed this finding, and the association of LASP1 with eIF4B via co-immunoprecipitation assays. Furthermore, we show that LASP1 can interact with eIF4A and eIF4B through a GST-pulldown approach. Activation of CXCR4 signaling increased the translation of oncoproteins downstream of eIF4A. Interestingly, genetic silencing of LASP1 interrupted the ability of eIF4A to translate oncogenic mRNAs into oncoproteins. This impaired ability of eIF4A was confirmed by a previously established 5'UTR luciferase reporter assay. Finally, lack of LASP1 sensitizes 231S cells to pharmacological inhibition of eIF4A by Rocaglamide A as evident through BIRC5 expression. Overall, our work identified the CXCR4-LASP1 axis to be a novel mediator in oncogenic protein translation. Thus, our axis of study represents a potential target for future TNBC therapies.
Keywords: CXCR4; LASP1; breast cancer; eIF4A1; eIF4B; eIF4F; protein translation.
Figures


Similar articles
-
The CXCR4-Dependent LASP1-Ago2 Interaction in Triple-Negative Breast Cancer.Cancers (Basel). 2020 Aug 29;12(9):2455. doi: 10.3390/cancers12092455. Cancers (Basel). 2020. PMID: 32872485 Free PMC article.
-
Role of the CXCR4-LASP1 Axis in the Stabilization of Snail1 in Triple-Negative Breast Cancer.Cancers (Basel). 2020 Aug 21;12(9):2372. doi: 10.3390/cancers12092372. Cancers (Basel). 2020. PMID: 32825729 Free PMC article.
-
Transcriptome-wide characterization of the eIF4A signature highlights plasticity in translation regulation.Genome Biol. 2014;15(10):476. doi: 10.1186/s13059-014-0476-1. Genome Biol. 2014. PMID: 25273840 Free PMC article.
-
LASP1 in Cellular Signaling and Gene Expression: More than Just a Cytoskeletal Regulator.Cells. 2022 Nov 29;11(23):3817. doi: 10.3390/cells11233817. Cells. 2022. PMID: 36497077 Free PMC article. Review.
-
An update on the LIM and SH3 domain protein 1 (LASP1): a versatile structural, signaling, and biomarker protein.Oncotarget. 2015 Jan 1;6(1):26-42. doi: 10.18632/oncotarget.3083. Oncotarget. 2015. PMID: 25622104 Free PMC article. Review.
Cited by
-
Targeting of the Eukaryotic Translation Initiation Factor 4A Against Breast Cancer Stemness.Front Oncol. 2019 Dec 6;9:1311. doi: 10.3389/fonc.2019.01311. eCollection 2019. Front Oncol. 2019. PMID: 31867270 Free PMC article.
-
Long noncoding RNA PPP1R14B-AS1 imitates microRNA-134-3p to facilitate breast cancer progression by upregulating LIM and SH3 protein 1.Oncol Res. 2022 Aug 31;29(4):251-262. doi: 10.32604/or.2022.03582. eCollection 2021. Oncol Res. 2022. PMID: 37303940 Free PMC article.
-
Nephrotoxicity Profile of Cadmium Revealed by Proteomics in Mouse Kidney.Biol Trace Elem Res. 2021 May;199(5):1929-1940. doi: 10.1007/s12011-020-02312-7. Epub 2020 Aug 15. Biol Trace Elem Res. 2021. PMID: 32803525
-
The CXCR4-Dependent LASP1-Ago2 Interaction in Triple-Negative Breast Cancer.Cancers (Basel). 2020 Aug 29;12(9):2455. doi: 10.3390/cancers12092455. Cancers (Basel). 2020. PMID: 32872485 Free PMC article.
-
Role of the CXCR4-LASP1 Axis in the Stabilization of Snail1 in Triple-Negative Breast Cancer.Cancers (Basel). 2020 Aug 21;12(9):2372. doi: 10.3390/cancers12092372. Cancers (Basel). 2020. PMID: 32825729 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

