A comprehensive search of functional sequence space using large mammalian display libraries created by gene editing
- PMID: 31107136
- PMCID: PMC6601556
- DOI: 10.1080/19420862.2019.1618673
A comprehensive search of functional sequence space using large mammalian display libraries created by gene editing
Abstract
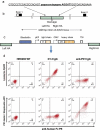
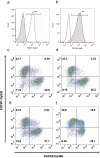
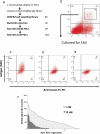
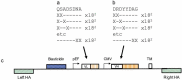
The construction of large libraries in mammalian cells allows the direct screening of millions of molecular variants for binding properties in a cell type relevant for screening or production. We have created mammalian cell libraries of up to 10 million clones displaying a repertoire of IgG-formatted antibodies on the cell surface. TALE nucleases or CRISPR/Cas9 were used to direct the integration of the antibody genes into a single genomic locus, thereby rapidly achieving stable expression and transcriptional normalization. The utility of the system is illustrated by the affinity maturation of a PD-1-blocking antibody through the systematic mutation and functional survey of 4-mer variants within a 16 amino acid paratope region. Mutating VH CDR3 only, we identified a dominant "solution" involving substitution of a central tyrosine to histidine. This appears to be a local affinity maximum, and this variant was surpassed by a lysine substitution when light chain variants were introduced. We achieve this comprehensive and quantitative interrogation of sequence space by combining high-throughput oligonucleotide synthesis with mammalian display and flow cytometry operating at the multi-million scale.
Keywords: CRISPR/Cas9; IgG antibody library; Mammalian display; TALE nuclease; affinity maturation; fluorescence-activated cell sorting; gene editing; gene targeting; human therapeutic antibody discovery; magnetic-activated cell sorting.
Figures
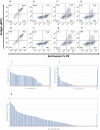


References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
