Cancer Cells Tune the Signaling Pathways to Empower de Novo Synthesis of Nucleotides
- PMID: 31108873
- PMCID: PMC6562601
- DOI: 10.3390/cancers11050688
Cancer Cells Tune the Signaling Pathways to Empower de Novo Synthesis of Nucleotides
Abstract
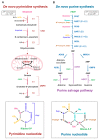
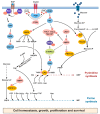
Cancer cells exhibit a dynamic metabolic landscape and require a sufficient supply of nucleotides and other macromolecules to grow and proliferate. To meet the metabolic requirements for cell growth, cancer cells must stimulate de novo nucleotide synthesis to obtain adequate nucleotide pools to support nucleic acid and protein synthesis along with energy preservation, signaling activity, glycosylation mechanisms, and cytoskeletal function. Both oncogenes and tumor suppressors have recently been identified as key molecular determinants for de novo nucleotide synthesis that contribute to the maintenance of homeostasis and the proliferation of cancer cells. Inactivation of tumor suppressors such as TP53 and LKB1 and hyperactivation of the mTOR pathway and of oncogenes such as MYC, RAS, and AKT have been shown to fuel nucleotide synthesis in tumor cells. The molecular mechanisms by which these signaling hubs influence metabolism, especially the metabolic pathways for nucleotide synthesis, continue to emerge. Here, we focus on the current understanding of the molecular mechanisms by which oncogenes and tumor suppressors modulate nucleotide synthesis in cancer cells and, based on these insights, discuss potential strategies to target cancer cell proliferation.
Keywords: AKT; MYC; RAS; cancer metabolism; de novo nucleotide synthesis; mTORC1; metabolic vulnerability; oncogenes; short term and long-term regulation; tumor suppressors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

