Chromosome Arm Locations of Barley Sucrose Transporter Gene in Transgenic Winter Wheat Lines
- PMID: 31114602
- PMCID: PMC6502970
- DOI: 10.3389/fpls.2019.00548
Chromosome Arm Locations of Barley Sucrose Transporter Gene in Transgenic Winter Wheat Lines
Abstract
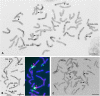
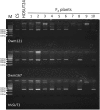
Three transgenic HOSUT lines of winter wheat, HOSUT12, HOSUT20, and HOSUT24, each harbor a single copy of the cDNA for the barley sucrose transporter gene HvSUT1 (SUT), which was fused to the barley endosperm-specific Hordein B1 promoter (HO; the HOSUT transgene). Previously, flow cytometry combined with PCR analysis demonstrated that the HOSUT transgene had been integrated into different wheat chromosomes: 7A, 5D, and 4A in HOSUT12, HOSUT20, and HOSUT24, respectively. In order to confirm the chromosomal location of the HOSUT transgene by a cytological approach using wheat aneuploid stocks, we crossed corresponding nullisomic-tetrasomic lines with the three HOSUT lines, namely nullisomic 7A-tetrasomic 7B with HOSUT12, nullisomic 5D-tetrasomic 5B with HOSUT20, and nullisomic 4A-tetrasomic 4B with HOSUT24. We examined the resulting chromosomal constitutions and the presence of the HOSUT transgene in the F2 progeny by means of chromosome banding and PCR. The chromosome banding patterns of the critical chromosomes in the original HOSUT lines showed no difference from those of the corresponding wild type chromosomes. The presence or absence of the critical chromosomes completely corresponded to the presence or absence of the HOSUT transgene in the F2 plants. Investigating telocentric chromosomes occurred in the F2 progeny, which were derived from the respective critical HOSUT chromosomes, we found that the HOSUT transgene was individually integrated on the long arms of chromosomes 4A, 7A, and 5D in the three HOSUT lines. Thus, in this study we verified the chromosomal locations of the transgene, which had previously been determined by flow cytometry, and moreover revealed the chromosome-arm locations of the HOSUT transgene in the HOSUT lines.
Keywords: HOSUT lines; barley sucrose transporter; chromosome banding; nullisomic-tetrasomics; telocentric chromosome; transgene; wheat.
Figures




References
-
- Cain-Hom C., Splinter E., van Min M., Simonis M., van de Heijning M., Martinez M., et al. (2017). Efficient mapping of transgene integration sites and local structural changes in Cre transgenic mice using targeted locus amplification. Nucleic Acids Res. 45:e62. 10.1093/nar/gkw1329 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

