Promoter-proximal pausing of RNA polymerase II: a nexus of gene regulation
- PMID: 31123063
- PMCID: PMC6672056
- DOI: 10.1101/gad.325142.119
Promoter-proximal pausing of RNA polymerase II: a nexus of gene regulation
Abstract
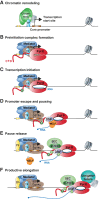
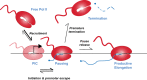
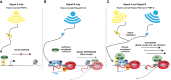
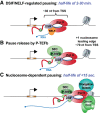
Precise spatio-temporal control of gene activity is essential for organismal development, growth, and survival in a changing environment. Decisive steps in gene regulation involve the pausing of RNA polymerase II (Pol II) in early elongation, and the controlled release of paused polymerase into productive RNA synthesis. Here we describe the factors that enable pausing and the events that trigger Pol II release into the gene. We also discuss open questions in the field concerning the stability of paused Pol II, nucleosomes as obstacles to elongation, and potential roles of pausing in defining the precision and dynamics of gene expression.
Keywords: Pol II pausing; gene expression; gene regulation; transcription regulation.
© 2019 Core and Adelman; Published by Cold Spring Harbor Laboratory Press.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
