In silico survey of the central conserved regions in viroids of the Pospiviroidae family for conserved asymmetric loop structures
- PMID: 31123078
- PMCID: PMC6633198
- DOI: 10.1261/rna.070409.119
In silico survey of the central conserved regions in viroids of the Pospiviroidae family for conserved asymmetric loop structures
Abstract
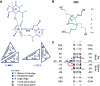
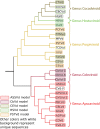
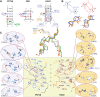
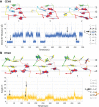
Viroids are the smallest replicative pathogens, consisting of RNA circles (∼300 nucleotides) that require host machinery to replicate. Structural RNA elements recruit these host factors. Currently, many of these structural elements and the nature of their interactions are unknown. All Pospiviroidae have homology in the central conserved region (CCR). The CCR of potato spindle tuber viroid (PSTVd) contains a sarcin/ricin domain (SRD), the only viroid structural element with an unequivocal replication role. We assumed that every member of this family uses this region to recruit host factors, and that each CCR has an SRD-like asymmetric loop within it. Potential SRD or SRD-like motifs were sought in the CCR of each Pospiviroidae member as follows. Motif location in each CCR was predicted with MUSCLE alignment and Vienna RNAfold. Viroid-specific models of SRD-like motifs were built by superimposing noncanonical base pairs and nucleotides on a model of an SRD. The RNA geometry search engine FR3D was then used to find nucleotide groups close to the geometry suggested by this superimposition. Atomic resolution structures were assembled using the molecular visualization program Chimera, and the stability of each motif was assessed with molecular dynamics (MD). Some models required a protonated cytosine. To be stable within a cell, the pKa of that cytosine must be shifted up. Constant pH-replica exchange MD analysis showed such a shift in the proposed structures. These data show that every Pospiviroidae member could form a motif that resembles an SRD in its CCR, and imply there could be undiscovered mimics of other RNA domains.
Keywords: Loop E RNA; RNA domains; RNA dynamics; RNA mimicry; S-turn; constant pH replica exchange molecular dynamics; molecular dynamics; sarcin/ricin domain.
© 2019 Freidhoff and Bruist; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures


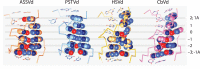


References
-
- Calladine C, Drew H, Luisi B, Travers A. 2004. Understanding DNA, 3rd ed Academic Press, MA.
-
- Case DA, Babin V, Berryman JT, Betz RM, Cai Q, Cerutti DS, Cheatham TE III, Darden TA, Duke RE, Gohlke H, et al. 2014. AMBER 14. University of California, San Francisco: http://ambermd.org/Manuals.php
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
