The open diffusion data derivatives, brain data upcycling via integrated publishing of derivatives and reproducible open cloud services
- PMID: 31123325
- PMCID: PMC6533280
- DOI: 10.1038/s41597-019-0073-y
The open diffusion data derivatives, brain data upcycling via integrated publishing of derivatives and reproducible open cloud services
Abstract
We describe the Open Diffusion Data Derivatives (O3D) repository: an integrated collection of preserved brain data derivatives and processing pipelines, published together using a single digital-object-identifier. The data derivatives were generated using modern diffusion-weighted magnetic resonance imaging data (dMRI) with diverse properties of resolution and signal-to-noise ratio. In addition to the data, we publish all processing pipelines (also referred to as open cloud services). The pipelines utilize modern methods for neuroimaging data processing (diffusion-signal modelling, fiber tracking, tractography evaluation, white matter segmentation, and structural connectome construction). The O3D open services can allow cognitive and clinical neuroscientists to run the connectome mapping algorithms on new, user-uploaded, data. Open source code implementing all O3D services is also provided to allow computational and computer scientists to reuse and extend the processing methods. Publishing both data-derivatives and integrated processing pipeline promotes practices for scientific reproducibility and data upcycling by providing open access to the research assets for utilization by multiple scientific communities.
Conflict of interest statement
The authors declare no competing interests.
Figures
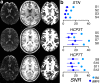
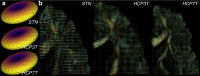
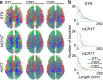

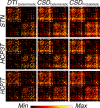
References
Publication types
MeSH terms
Grants and funding
- P30 AG053760/AG/NIA NIH HHS/United States
- U01 MH097435/MH/NIMH NIH HHS/United States
- R01 AG019771/AG/NIA NIH HHS/United States
- ULTTR001108/U.S. Department of Health & Human Services | NIH | National Institute on Minority Health and Health Disparities (NIMHD)/International
- U54 EB020406/EB/NIBIB NIH HHS/United States
- 1445604/NSF | Directorate for Computer and Information Science and Engineering (Directorate for Computer & Information Science & Engineering)/International
- P30 AG010133/AG/NIA NIH HHS/United States
- 1636893/NSF | Directorate for Computer and Information Science and Engineering (Directorate for Computer & Information Science & Engineering)/International
- U01 AG024904/AG/NIA NIH HHS/United States
- T32 MH103213/MH/NIMH NIH HHS/United States
- P20 NR015331/NR/NINR NIH HHS/United States
- P30 DK089503/DK/NIDDK NIH HHS/United States
- R01 LM011360/LM/NLM NIH HHS/United States
- U54 MH091657/MH/NIMH NIH HHS/United States
- R01 CA129769/CA/NCI NIH HHS/United States
- P50 NS091856/NS/NINDS NIH HHS/United States
- 1734853/NSF | Directorate for Social, Behavioral & Economic Sciences | Division of Behavioral and Cognitive Sciences (Behavioral & Cognitive Sciences)/International
LinkOut - more resources
Full Text Sources

