Quantitative Proteomic Analysis Reveals Novel Insights into Intracellular Silicate Stress-Responsive Mechanisms in the Diatom Skeletonema dohrnii
- PMID: 31126124
- PMCID: PMC6566588
- DOI: 10.3390/ijms20102540
Quantitative Proteomic Analysis Reveals Novel Insights into Intracellular Silicate Stress-Responsive Mechanisms in the Diatom Skeletonema dohrnii
Abstract
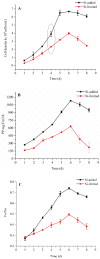
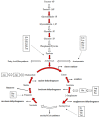
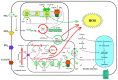
Diatoms are a successful group of marine phytoplankton that often thrives under adverse environmental stress conditions. Members of the Skeletonema genus are ecologically important which may subsist during silicate stress and form a dense bloom following higher silicate concentration. However, our understanding of diatoms' underlying molecular mechanism involved in these intracellular silicate stress-responses are limited. Here an iTRAQ-based proteomic method was coupled with multiple physiological techniques to explore distinct cellular responses associated with oxidative stress in the diatom Skeletonema dohrnii to the silicate limitation. In total, 1768 proteins were detected; 594 proteins were identified as differentially expressed (greater than a two-fold change; p < 0.05). In Si-limited cells, downregulated proteins were mainly related to photosynthesis metabolism, light-harvesting complex, and oxidative phosphorylation, corresponding to inducing oxidative stress, and ROS accumulation. None of these responses were identified in Si-limited cells; in comparing with other literature, Si-stress cells showed that ATP-limited diatoms are unable to rely on photosynthesis, which will break down and reshuffle carbon metabolism to compensate for photosynthetic carbon fixation losses. Our findings have a good correlation with earlier reports and provides a new molecular level insight into the systematic intracellular responses employed by diatoms in response to silicate stress in the marine environment.
Keywords: abiotic stress; carbon fixation; diatom; iTRAQ; photosynthesis; proteomics; silicate limitation.
Conflict of interest statement
The authors declare no conflict of interest
Figures



Similar articles
-
Quantitative Proteomic Profiling of Marine Diatom Skeletonema dohrnii in Response to Temperature and Silicate Induced Environmental Stress.Front Microbiol. 2021 Jan 14;11:554832. doi: 10.3389/fmicb.2020.554832. eCollection 2020. Front Microbiol. 2021. PMID: 33519723 Free PMC article.
-
Comparative Proteomic Analysis Reveals New Insights Into the Common and Specific Metabolic Regulation of the Diatom Skeletonema dohrnii to the Silicate and Temperature Availability.Front Plant Sci. 2020 Nov 5;11:578915. doi: 10.3389/fpls.2020.578915. eCollection 2020. Front Plant Sci. 2020. PMID: 33224167 Free PMC article.
-
Diatom carbon export enhanced by silicate upwelling in the northeast Atlantic.Nature. 2005 Sep 29;437(7059):728-32. doi: 10.1038/nature03948. Nature. 2005. PMID: 16193051
-
Proteomics studies on stress responses in diatoms.Proteomics. 2015 Dec;15(23-24):3943-53. doi: 10.1002/pmic.201500165. Epub 2015 Oct 12. Proteomics. 2015. PMID: 26364674 Review.
-
Inorganic carbon availability in benthic diatom communities: photosynthesis and migration.Philos Trans R Soc Lond B Biol Sci. 2017 Sep 5;372(1728):20160398. doi: 10.1098/rstb.2016.0398. Philos Trans R Soc Lond B Biol Sci. 2017. PMID: 28717024 Free PMC article. Review.
Cited by
-
The Biotechnological Potential of the Marine Diatom Skeletonema dohrnii to the Elevated Temperature and pCO2 Concentration.Mar Drugs. 2020 May 15;18(5):259. doi: 10.3390/md18050259. Mar Drugs. 2020. PMID: 32429035 Free PMC article.
-
Protein and Proteome Atlas for Plants under Stresses: New Highlights and Ways for Integrated Omics in Post-Genomics Era.Int J Mol Sci. 2019 Oct 21;20(20):5222. doi: 10.3390/ijms20205222. Int J Mol Sci. 2019. PMID: 31640274 Free PMC article. Review.
-
Quantitative Proteomic Profiling of Marine Diatom Skeletonema dohrnii in Response to Temperature and Silicate Induced Environmental Stress.Front Microbiol. 2021 Jan 14;11:554832. doi: 10.3389/fmicb.2020.554832. eCollection 2020. Front Microbiol. 2021. PMID: 33519723 Free PMC article.
-
Comparative Proteomic Analysis Reveals New Insights Into the Common and Specific Metabolic Regulation of the Diatom Skeletonema dohrnii to the Silicate and Temperature Availability.Front Plant Sci. 2020 Nov 5;11:578915. doi: 10.3389/fpls.2020.578915. eCollection 2020. Front Plant Sci. 2020. PMID: 33224167 Free PMC article.
References
-
- Rosenwasser S., Van Creveld S.G., Schatz D., Malitsky S., Tzfadia O., Aharoni A., Levin Y., Gabashvili A., Feldmesser E., Vardi A. Mapping the Diatom Redox-Sensitive Proteome Provides Insight into Response to Nitrogen Stress in the Marine Environment. Proc. Natl. Acad. Sci. USA. 2014;111:2740–2745. doi: 10.1073/pnas.1319773111. - DOI - PMC - PubMed
-
- Du C., Liang J.-R., Chen D.-D., Xu B., Zhuo W.-H., Gao Y.-H., Chen C.-P., Bowler C., Zhang W. iTRAQ-Based Proteomic Analysis of the Metabolism Mechanism Associated with Silicon Response in the Marine Diatom Thalassiosira Pseudonana. J. Proteome Res. 2014;13:720–734. doi: 10.1021/pr400803w. - DOI - PubMed
-
- Brzezinski M.A., Olson R.J., Chisholm S.W. Silicon Availability and Cell-Cycle Progression in Marine Diatoms. Mar. Ecol. Prog. Ser. 1990;67:83–96. doi: 10.3354/meps067083. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

