Gypsum-DL: an open-source program for preparing small-molecule libraries for structure-based virtual screening
- PMID: 31127411
- PMCID: PMC6534830
- DOI: 10.1186/s13321-019-0358-3
Gypsum-DL: an open-source program for preparing small-molecule libraries for structure-based virtual screening
Abstract
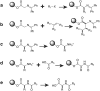
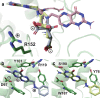
Computational techniques such as structure-based virtual screening require carefully prepared 3D models of potential small-molecule ligands. Though powerful, existing commercial programs for virtual-library preparation have restrictive and/or expensive licenses. Freely available alternatives, though often effective, do not fully account for all possible ionization, tautomeric, and ring-conformational variants. We here present Gypsum-DL, a free, robust open-source program that addresses these challenges. As input, Gypsum-DL accepts virtual compound libraries in SMILES or flat SDF formats. For each molecule in the virtual library, it enumerates appropriate ionization, tautomeric, chiral, cis/trans isomeric, and ring-conformational forms. As output, Gypsum-DL produces an SDF file containing each molecular form, with 3D coordinates assigned. To demonstrate its utility, we processed 1558 molecules taken from the NCI Diversity Set VI and 56,608 molecules taken from a Distributed Drug Discovery (D3) combinatorial virtual library. We also used 4463 high-quality protein-ligand complexes from the PDBBind database to show that Gypsum-DL processing can improve virtual-screening pose prediction. Gypsum-DL is available free of charge under the terms of the Apache License, Version 2.0.
Keywords: 3D structure generation; Chemical biology; Computational biology; Computer-aided drug discovery; Small-molecule libraries; Virtual screening.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

