A totally data-driven whole-brain multimodal pipeline for the discrimination of Parkinson's disease, multiple system atrophy and healthy control
- PMID: 31128523
- PMCID: PMC6531871
- DOI: 10.1016/j.nicl.2019.101858
A totally data-driven whole-brain multimodal pipeline for the discrimination of Parkinson's disease, multiple system atrophy and healthy control
Abstract
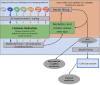
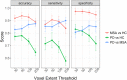
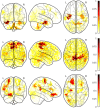
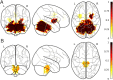
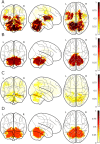
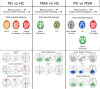
Parkinson's Disease (PD) and Multiple System Atrophy (MSA) are two parkinsonian syndromes that share many symptoms, albeit having very different prognosis. Although previous studies have proposed multimodal MRI protocols combined with multivariate analysis to discriminate between these two populations and healthy controls, studies combining all MRI indexes relevant for these disorders (i.e. grey matter volume, fractional anisotropy, mean diffusivity, iron deposition, brain activity at rest and brain connectivity) with a completely data-driven voxelwise analysis for discrimination are still lacking. In this study, we used such a complete MRI protocol and adapted a fully-data driven analysis pipeline to discriminate between these populations and a healthy controls (HC) group. The pipeline combined several feature selection and reduction steps to obtain interpretable models with a low number of discriminant features that can shed light onto the brain pathology of PD and MSA. Using this pipeline, we could discriminate between PD and HC (best accuracy = 0.78), MSA and HC (best accuracy = 0.94) and PD and MSA (best accuracy = 0.88). Moreover, we showed that indexes derived from resting-state fMRI alone could discriminate between PD and HC, while mean diffusivity in the cerebellum and the putamen alone could discriminate between MSA and HC. On the other hand, a more diverse set of indexes derived by multiple modalities was needed to discriminate between the two disorders. We showed that our pipeline was able to discriminate between distinct pathological populations while delivering sparse model that could be used to better understand the neural underpinning of the pathologies.
Keywords: Data-driven clinical classification; Multimodal MRI; Parkinsonism discrimination.
Copyright © 2019 The Authors. Published by Elsevier Inc. All rights reserved.
Figures



Similar articles
-
Cerebellar resting-state functional connectivity in Parkinson's disease and multiple system atrophy: Characterization of abnormalities and potential for differential diagnosis at the single-patient level.Neuroimage Clin. 2019;22:101720. doi: 10.1016/j.nicl.2019.101720. Epub 2019 Feb 13. Neuroimage Clin. 2019. PMID: 30785051 Free PMC article.
-
MRI supervised and unsupervised classification of Parkinson's disease and multiple system atrophy.Mov Disord. 2018 Apr;33(4):600-608. doi: 10.1002/mds.27307. Epub 2018 Feb 23. Mov Disord. 2018. PMID: 29473662
-
Diagnostic Potential of Multimodal MRI Markers in Atypical Parkinsonian Disorders.J Parkinsons Dis. 2019;9(4):681-691. doi: 10.3233/JPD-181568. J Parkinsons Dis. 2019. PMID: 31450511
-
Voxelwise meta-analysis of gray matter anomalies in Parkinson variant of multiple system atrophy and Parkinson's disease using anatomic likelihood estimation.Neurosci Lett. 2015 Feb 5;587:79-86. doi: 10.1016/j.neulet.2014.12.007. Epub 2014 Dec 5. Neurosci Lett. 2015. PMID: 25484255
-
The role of functional neuroimaging in the differential diagnosis of idiopathic Parkinson's disease and multiple system atrophy.Clin Auton Res. 2004 Apr;14(2):84-91. doi: 10.1007/s10286-004-0167-1. Clin Auton Res. 2004. PMID: 15095050 Review.
Cited by
-
Reproducible metabolic topographies associated with multiple system atrophy: Network and regional analyses in Chinese and American patient cohorts.Neuroimage Clin. 2020;28:102416. doi: 10.1016/j.nicl.2020.102416. Epub 2020 Sep 9. Neuroimage Clin. 2020. PMID: 32987300 Free PMC article.
-
Multimodal striatal neuromarkers in distinguishing parkinsonian variant of multiple system atrophy from idiopathic Parkinson's disease.CNS Neurosci Ther. 2022 Dec;28(12):2172-2182. doi: 10.1111/cns.13959. Epub 2022 Sep 1. CNS Neurosci Ther. 2022. PMID: 36047435 Free PMC article.
-
Automated Analysis of Diffusion-Weighted Magnetic Resonance Imaging for the Differential Diagnosis of Multiple System Atrophy from Parkinson's Disease.Mov Disord. 2021 Jan;36(1):241-245. doi: 10.1002/mds.28281. Epub 2020 Sep 16. Mov Disord. 2021. PMID: 32935402 Free PMC article.
-
Proportional Changes in Cognitive Subdomains During Normal Brain Aging.Front Aging Neurosci. 2021 Nov 15;13:673469. doi: 10.3389/fnagi.2021.673469. eCollection 2021. Front Aging Neurosci. 2021. PMID: 34867263 Free PMC article.
-
Brain-age estimation accuracy is significantly increased using multishell free-water reconstruction.Hum Brain Mapp. 2022 May;43(7):2365-2376. doi: 10.1002/hbm.25792. Epub 2022 Feb 10. Hum Brain Mapp. 2022. PMID: 35141974 Free PMC article.
References
-
- Anderson J., Jenkinson M., Smith N. Non-linear registration, aka spatial normalisation. 2010. http://www.fmrib.ox.ac.uk/datasets/techrep/ Retrieved from.
-
- Arlicot N., Vercouillie J., Ribeiro M.-J., Tauber C., Venel Y., Baulieu J.-L., Guilloteau D. Initial evaluation in healthy humans of [18F]DPA-714, a potential PET biomarker for neuroinflammation. Nucl. Med. Biol. 2012;39(4):570–578. - PubMed
-
- Ashburner J. A fast diffeomorphic image registration algorithm. NeuroImage. 2007;38(1):95–113. - PubMed
-
- Baddeley A., Rubak E., Turner R. Spatial point patterns : methodology and applications with R. 2015. http://spatstat.org/ Retrieved from.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

