5',8-Cyclopurine Lesions in DNA Damage: Chemical, Analytical, Biological, and Diagnostic Significance
- PMID: 31141888
- PMCID: PMC6628319
- DOI: 10.3390/cells8060513
5',8-Cyclopurine Lesions in DNA Damage: Chemical, Analytical, Biological, and Diagnostic Significance
Abstract
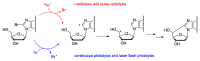
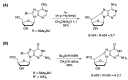
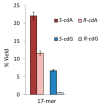
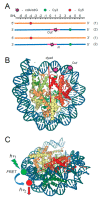
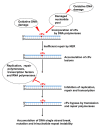
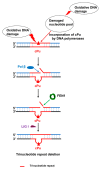
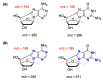
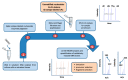
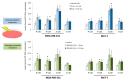
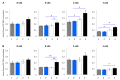
Purine 5',8-cyclo-2'-deoxynucleosides (cPu) are tandem-type lesions observed among the DNA purine modifications and identified in mammalian cellular DNA in vivo. These lesions can be present in two diasteroisomeric forms, 5'R and 5'S, for each 2'-deoxyadenosine and 2'-deoxyguanosine moiety. They are generated exclusively by hydroxyl radical attack to 2'-deoxyribose units generating C5' radicals, followed by cyclization with the C8 position of the purine base. This review describes the main recent achievements in the preparation of the cPu molecular library for analytical and DNA synthesis applications for the studies of the enzymatic recognition and repair mechanisms, their impact on transcription and genetic instability, quantitative determination of the levels of lesions in various types of cells and animal model systems, and relationships between the levels of lesions and human health, disease, and aging, as well as the defining of the detection limits and quantification protocols.
Keywords: DNA and RNA polymerases; DNA damage; LC-MS/MS; cancer; cyclopurines; free radicals; nucleotide excision repair; reactive oxygen species; xeroderma pigmentosum.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
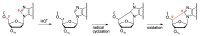


Similar articles
-
Mass spectrometric assays for the tandem lesion 8,5'-cyclo-2'-deoxyguanosine in mammalian DNA.Biochemistry. 2002 Mar 19;41(11):3703-11. doi: 10.1021/bi016004d. Biochemistry. 2002. PMID: 11888287
-
Synthesis and characterization of oligonucleotides containing 5',8-cyclopurine 2'-deoxyribonucleosides: (5'R)-5',8-cyclo-2'-deoxyadenosine, (5'S)-5',8-cyclo-2'-deoxyguanosine, and (5'R)-5',8-cyclo-2'-deoxyguanosine.Chem Res Toxicol. 1999 May;12(5):412-21. doi: 10.1021/tx9802668. Chem Res Toxicol. 1999. PMID: 10328751
-
Purine DNA Lesions at Different Oxygen Concentration in DNA Repair-Impaired Human Cells (EUE-siXPA).Cells. 2019 Nov 1;8(11):1377. doi: 10.3390/cells8111377. Cells. 2019. PMID: 31683970 Free PMC article.
-
Purine 5',8-cyclonucleoside lesions: chemistry and biology.Chem Soc Rev. 2011 Mar;40(3):1368-82. doi: 10.1039/c0cs00061b. Epub 2011 Jan 11. Chem Soc Rev. 2011. PMID: 21221459 Review.
-
On the relevance of hydroxyl radical to purine DNA damage.Free Radic Res. 2021 Apr;55(4):384-404. doi: 10.1080/10715762.2021.1876855. Epub 2021 Jan 26. Free Radic Res. 2021. PMID: 33494618 Review.
Cited by
-
Effects of 5',8'-Cyclo-2'-Deoxypurines on the Base Excision Repair of Clustered DNA Lesions in Nuclear Extracts of the XPC Cell Line.Cells. 2021 Nov 20;10(11):3254. doi: 10.3390/cells10113254. Cells. 2021. PMID: 34831476 Free PMC article.
-
Ne-22 Ion-Beam Radiation Damage to DNA: From Initial Free Radical Formation to Resulting DNA-Base Damage.ACS Omega. 2021 Jun 14;6(25):16600-16611. doi: 10.1021/acsomega.1c01954. eCollection 2021 Jun 29. ACS Omega. 2021. PMID: 34235332 Free PMC article.
-
An overview of current advances in perinatal alcohol exposure and pathogenesis of fetal alcohol spectrum disorders.J Neurodev Disord. 2024 Apr 20;16(1):20. doi: 10.1186/s11689-024-09537-w. J Neurodev Disord. 2024. PMID: 38643092 Free PMC article. Review.
-
Advances in Nucleic Acid Research: Exploring the Potential of Oligonucleotides for Therapeutic Applications and Biological Studies.Int J Mol Sci. 2023 Dec 21;25(1):146. doi: 10.3390/ijms25010146. Int J Mol Sci. 2023. PMID: 38203317 Free PMC article. Review.
-
New Insights into the Reaction Paths of Hydroxyl Radicals with Purine Moieties in DNA and Double-Stranded Oligodeoxynucleotides.Molecules. 2019 Oct 26;24(21):3860. doi: 10.3390/molecules24213860. Molecules. 2019. PMID: 31717733 Free PMC article.
References
-
- McCord J.M., Fridovich I. The reduction of cytochrome c by milk xanthine oxidase. J. Biol. Chem. 1968;243:5753–5760. - PubMed
-
- Winterbourn C.C. Biological chemistry of reactive oxygen species. In: Chatgilialoglu C., Studer A., editors. Encyclopedia of Radicals in Chemistry, Biology and Materials. Volume 3. Wiley; Chichester, UK: 2012. pp. 1260–1281.
-
- Geacintov N.E., Shafirovich V. Reactions of small reactive species with DNA. In: Chatgilialoglu C., Studer A., editors. Encyclopedia of Radicals in Chemistry, Biology and Materials. Volume 3. Wiley; Chichester, UK: 2012. pp. 1284–1317.
-
- Friedberg E.C., Walker G.C., Siede W. DNA Repair Mutagenesis. ASM Press; Washington, WA, USA: 1995.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

